GenomeSequence Class Reference
Create/Access/Modify/Load Genome Sequences stored as binary mapped files. More...
#include <GenomeSequence.h>
Public Member Functions | |
| GenomeSequence () | |
| Simple constructor - no implicit file open. | |
| void | constructorClear () |
| GenomeSequence (std::string &referenceFilename) | |
| attempt to open an existing sequence | |
| GenomeSequence (const char *referenceFilename) | |
| Smarter constructor - attempt to open an existing sequence. | |
| ~GenomeSequence () | |
| Close the file if open and destroy the object. | |
| bool | open (bool isColorSpace=false, int flags=O_RDONLY) |
| open the reference specified using GenomeSequence::setReferenceName | |
| bool | open (const char *filename, int flags=O_RDONLY) |
| open the given file as the genome (no filename munging occurs). | |
| bool | create () |
| bool | create (bool isColor) |
| void | setProgressStream (std::ostream &progressStream) |
| void | setColorSpace (bool colorSpace) |
| void | setSearchCommonFileSuffix (bool searchCommonFileSuffix) |
| bool | loadFastaData (const char *filename) |
| bool | setReferenceName (std::string referenceFilename) |
| set the reference name that will be used in open() | |
| void | setApplication (std::string application) |
| set the application name in the binary file header | |
| const std::string & | getFastaName () const |
| const std::string & | getReferenceName () const |
| bool | isColorSpace () const |
| tell us if we are a color space reference or not | |
| genomeIndex_t | getNumberBases () const |
| return the number of bases represented in this reference | |
| int | getChromosome (genomeIndex_t position) const |
| given a whole genome index, get the chromosome it is located in | |
| int | getChromosome (const char *chromosomeName) const |
| given a chromosome name, return the chromosome index | |
| int | getChromosomeCount () const |
| Return the number of chromosomes in the genome. | |
| genomeIndex_t | getChromosomeStart (int chromosomeIndex) const |
| given a chromosome, return the genome base it starts in | |
| genomeIndex_t | getChromosomeSize (int chromosomeIndex) const |
| given a chromosome, return its size in bases | |
| genomeIndex_t | getGenomePosition (const char *chromosomeName, unsigned int chromosomeIndex) const |
| given a chromosome name and position, return the genome position | |
| genomeIndex_t | getGenomePosition (int chromosome, unsigned int chromosomeIndex) const |
| given a chromosome index and position, return the genome position | |
| genomeIndex_t | getGenomePosition (const char *chromosomeName) const |
| given the chromosome name, get the corresponding 0 based genome index for the start of that chromosome | |
| genomeIndex_t | getGenomePosition (int chromosomeIndex) const |
| const std::string & | getBaseFilename () const |
| const char * | getChromosomeName (int chromosomeIndex) const |
| void | setDebugFlag (bool d) |
| genomeIndex_t | sequenceLength () const |
| const char * | chromosomeName (int chr) const |
| void | sanityCheck (MemoryMap &fasta) const |
| std::string | IntegerToSeq (unsigned int n, unsigned int wordsize) const |
| bool | wordMatch (unsigned int index, std::string &word) const |
| bool | printNearbyWords (unsigned int index, unsigned int variance, std::string &word) const |
| char | BasePair (char c) const |
| void | dumpSequenceSAMDictionary (std::ostream &) const |
| void | dumpHeaderTSV (std::ostream &) const |
| char | operator[] (genomeIndex_t index) const |
| Return the bases in base space or color space for within range index, ot. | |
| uint8_t | getInteger (genomeIndex_t index) const |
| void | set (genomeIndex_t index, char value) |
| uint8_t * | getDataPtr (genomeIndex_t index) |
| obtain the pointer to the raw data for other access methods | |
| void | getReverseRead (std::string &read) |
| void | getReverseRead (String &read) |
| int | debugPrintReadValidation (std::string &read, std::string &quality, char direction, genomeIndex_t readLocation, int sumQuality, int mismatchCount, bool recurse=true) |
| void | getString (std::string &str, int chromosome, genomeIndex_t index, int baseCount) const |
| void | getString (String &str, int chromosome, genomeIndex_t index, int baseCount) const |
| void | getString (std::string &str, genomeIndex_t index, int baseCount) const |
| void | getString (String &str, genomeIndex_t index, int baseCount) const |
| void | getHighLightedString (std::string &str, genomeIndex_t index, int baseCount, genomeIndex_t highLightStart, genomeIndex_t highLightEnd) const |
| void | print30 (genomeIndex_t) const |
| genomeIndex_t | simpleLocalAligner (std::string &read, std::string &quality, genomeIndex_t index, int windowSize) const |
| int | getMismatchCount (std::string &read, genomeIndex_t location, char exclude='\0') const |
| Return the mismatch count, disregarding CIGAR strings. | |
| int | getSumQ (std::string &read, std::string &qualities, genomeIndex_t location) const |
| brute force sumQ - no sanity checking | |
| void | getMismatchHatString (std::string &result, const std::string &read, genomeIndex_t location) const |
| void | getMismatchString (std::string &result, const std::string &read, genomeIndex_t location) const |
| void | getChromosomeAndIndex (std::string &, genomeIndex_t) const |
| void | getChromosomeAndIndex (String &, genomeIndex_t) const |
| bool | checkRead (std::string &read, std::string &qualities, std::string &cigar, int &sumQ, int &gapOpenCount, int &gapExtendCount, int &gapDeleteCount, std::string &result) const |
| check a SAM format read, using phred quality scores and the CIGAR string to determine if it is correct. | |
| bool | populateDBSNP (mmapArrayBool_t &dbSNP, std::ifstream &inputFile) const |
| bool | loadDBSNP (mmapArrayBool_t &dbSNP, const char *inputFileName) const |
| user friendly dbSNP loader. | |
Static Public Attributes | |
| static const int | baseAIndex = 000 |
| static const int | baseTIndex = 001 |
| static const int | baseCIndex = 002 |
| static const int | baseGIndex = 003 |
| static const int | baseNIndex = 004 |
| static const int | baseXIndex = 005 |
| static unsigned char | base2int [] |
| static const char | int2base [] = "ACGTNMXXXXXXXXXX" |
| static const char | int2colorSpace [] = "0123NXXXXXXXXXXX" |
| static unsigned char | base2complement [] |
Detailed Description
Create/Access/Modify/Load Genome Sequences stored as binary mapped files.
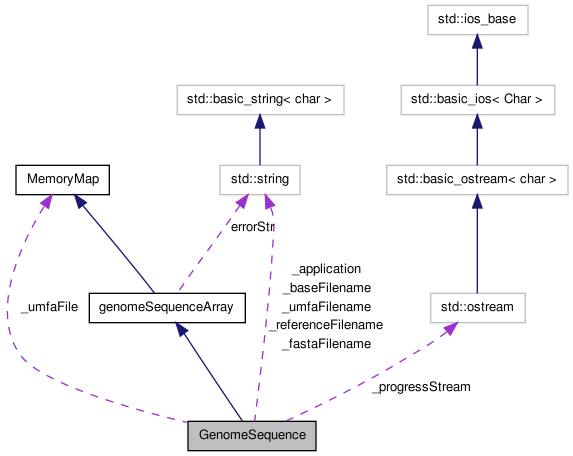
GenomeSequence is designed to be a high performance shared access reference object.
It is implemented as a MemoryMapArray template object with unsigned 8 bit ints, each of which stores two bases. Although 2 bits could be used, most references have more than four symbols (usually at least including 'N', indicating an unknown or masked out base).
Normal use of this class follows these steps:
- create the reference
- instantiate the GenomeSequence class object
- create the actual file (memory mapped) that is to hold the data
- populate the data using GenomeSequence::set
- use the reference
- use the reference by instantiating a GenomeSequence object
- either use the constructor with the reference filename
- or use GenomeSequence::setReferenceName() followed by open
- access the bases via the overloaded array operator []
- check sequence length by using GenomeSequence::getNumberBases()
- accessing chromosomes in the reference
- you typically will need to know about the chromosomes in the sequence
- see methods and docs with prefix 'getChromosome'
Sharing is accomplished using the mmap() function via the MemoryMap base class. This allows a potentially large genome reference to be shared among a number of simultaneously executing instances of one or more programs sharing the same reference.
Definition at line 238 of file GenomeSequence.h.
Constructor & Destructor Documentation
| GenomeSequence::GenomeSequence | ( | std::string & | referenceFilename | ) | [inline] |
attempt to open an existing sequence
- Parameters:
-
referenceFilename the name of the reference fasta file to open debug if true, additional debug information is printed
Definition at line 284 of file GenomeSequence.h.
| GenomeSequence::GenomeSequence | ( | const char * | referenceFilename | ) | [inline] |
Smarter constructor - attempt to open an existing sequence.
- Parameters:
-
referenceFilename the name of the reference fasta file to open debug if true, additional debug information is printed
Definition at line 294 of file GenomeSequence.h.
Member Function Documentation
| bool GenomeSequence::checkRead | ( | std::string & | read, | |
| std::string & | qualities, | |||
| std::string & | cigar, | |||
| int & | sumQ, | |||
| int & | gapOpenCount, | |||
| int & | gapExtendCount, | |||
| int & | gapDeleteCount, | |||
| std::string & | result | |||
| ) | const |
check a SAM format read, using phred quality scores and the CIGAR string to determine if it is correct.
- Parameters:
-
read the read in base space qualities the phred encoded qualities (Sanger, not Illumina) cigar the SAM file CIGAR column sumQ if >0 on entry, is checked against the computed sumQ insertions count of insertions found in
| int GenomeSequence::getChromosome | ( | const char * | chromosomeName | ) | const |
given a chromosome name, return the chromosome index
This is done via a linear search of the chromosome table in the header of the mapped file, so it is O(N)
- Parameters:
-
chromosomeName the name of the chromosome - exact match only
- Returns:
- 0-based index into chromosome table - INVALID_CHROMOSOME_INDEX if error
Definition at line 796 of file GenomeSequence.cpp.
| int GenomeSequence::getChromosome | ( | genomeIndex_t | position | ) | const |
given a whole genome index, get the chromosome it is located in
This is done via a binary search of the chromosome table in the header of the mapped file, so it is O(log(N))
- Parameters:
-
0-based position the base in the genome
- Returns:
- 0-based index into chromosome table - INVALID_CHROMOSOME_INDEX if error
Definition at line 719 of file GenomeSequence.cpp.
Referenced by getGenomePosition().
00720 { 00721 if (position == INVALID_GENOME_INDEX) return INVALID_CHROMOSOME_INDEX; 00722 00723 if (header->_chromosomeCount == 0) 00724 return INVALID_CHROMOSOME_INDEX; 00725 00726 int start = 0; 00727 int stop = header->_chromosomeCount - 1; 00728 00729 // eliminate case where position is in the last chromosome, since the loop 00730 // below falls off the end of the list if it in the last one. 00731 00732 if (position > header->_chromosomes[stop].start) 00733 return (stop); 00734 00735 while (start <= stop) 00736 { 00737 int middle = (start + stop) / 2; 00738 00739 if (position >= header->_chromosomes[middle].start && position < header->_chromosomes[middle + 1].start) 00740 return middle; 00741 00742 if (position == header->_chromosomes[middle + 1].start) 00743 return (middle + 1); 00744 00745 if (position > header->_chromosomes[middle + 1].start) 00746 start = middle + 1; 00747 00748 if (position < header->_chromosomes[middle].start) 00749 stop = middle - 1; 00750 } 00751 00752 return -1; 00753 }
| int GenomeSequence::getChromosomeCount | ( | ) | const |
Return the number of chromosomes in the genome.
- Returns:
- number of chromosomes in the genome
Definition at line 713 of file GenomeSequence.cpp.
| genomeIndex_t GenomeSequence::getChromosomeSize | ( | int | chromosomeIndex | ) | const [inline] |
given a chromosome, return its size in bases
- Parameters:
-
0-based chromosome index
- Returns:
- size of the chromosome in bases
Definition at line 411 of file GenomeSequence.h.
| genomeIndex_t GenomeSequence::getChromosomeStart | ( | int | chromosomeIndex | ) | const [inline] |
given a chromosome, return the genome base it starts in
- Parameters:
-
0-based chromosome index
- Returns:
- 0-based genome index of the base that starts the chromosome
Definition at line 401 of file GenomeSequence.h.
| uint8_t* GenomeSequence::getDataPtr | ( | genomeIndex_t | index | ) | [inline] |
obtain the pointer to the raw data for other access methods
this is a fairly ugly hack to reach into the raw genome vector, get the byte that encodes two bases, and return it. This is used by karma ReadIndexer::getSumQ to compare genome matchines by byte (two bases at a time) to speed it up.
Definition at line 557 of file GenomeSequence.h.
| genomeIndex_t GenomeSequence::getGenomePosition | ( | int | chromosome, | |
| unsigned int | chromosomeIndex | |||
| ) | const |
given a chromosome index and position, return the genome position
- Parameters:
-
chromosome index of the chromosome chromosomeIndex 1-based chromosome position
- Returns:
- genome index of the above chromosome position
Definition at line 770 of file GenomeSequence.cpp.
| genomeIndex_t GenomeSequence::getGenomePosition | ( | const char * | chromosomeName, | |
| unsigned int | chromosomeIndex | |||
| ) | const |
given a chromosome name and position, return the genome position
- Parameters:
-
chromosomeName name of the chromosome - exact match only chromosomeIndex 1-based chromosome position
- Returns:
- genome index of the above chromosome position
Definition at line 761 of file GenomeSequence.cpp.
Referenced by SamTags::createMDTag(), SamQuerySeqWithRefIter::reset(), SamQuerySeqWithRef::seqWithEquals(), and SamQuerySeqWithRef::seqWithoutEquals().
00764 { 00765 genomeIndex_t i = getGenomePosition(chromosomeName); 00766 if (i == INVALID_GENOME_INDEX) return INVALID_GENOME_INDEX; 00767 return i + chromosomeIndex - 1; 00768 }
| int GenomeSequence::getMismatchCount | ( | std::string & | read, | |
| genomeIndex_t | location, | |||
| char | exclude = '\0' | |||
| ) | const [inline] |
Return the mismatch count, disregarding CIGAR strings.
- Parameters:
-
read is the sequence we're counting mismatches in location is where in the genmoe we start comparing exclude is a wildcard character (e.g. '.' or 'N')
- Returns:
- number of bases that don't match the reference, except those that match exclude
Definition at line 619 of file GenomeSequence.h.
| genomeIndex_t GenomeSequence::getNumberBases | ( | ) | const [inline] |
return the number of bases represented in this reference
- Returns:
- count of bases
Definition at line 371 of file GenomeSequence.h.
Referenced by loadDBSNP(), and operator[]().
| int GenomeSequence::getSumQ | ( | std::string & | read, | |
| std::string & | qualities, | |||
| genomeIndex_t | location | |||
| ) | const [inline] |
brute force sumQ - no sanity checking
- Parameters:
-
read shotgun sequencer read string qualities phred quality string of same length location the alignment location to check sumQ
Definition at line 632 of file GenomeSequence.h.
| bool GenomeSequence::isColorSpace | ( | ) | const [inline] |
tell us if we are a color space reference or not
- Returns:
- true if colorspace, false otherwise
Definition at line 364 of file GenomeSequence.h.
Referenced by operator[]().
| bool GenomeSequence::loadDBSNP | ( | mmapArrayBool_t & | dbSNP, | |
| const char * | inputFileName | |||
| ) | const |
user friendly dbSNP loader.
- Parameters:
-
inputFileName may be empty, point to a text file or a dbSNP vector file
In all cases, dbSNP is returned the same length as this genome.
When no SNPs are loaded, all values are false.
When a text file is given, the file is parsed with two space separated columns - the first column is the chromosome name, and the second is the 1-based chromosome position of the SNP.
- Returns:
- false if a dbSNP file was correctly loaded, true otherwise
Definition at line 1252 of file GenomeSequence.cpp.
References MemoryMapArray< elementT, indexT, cookieVal, versionVal, accessorFunc, setterFunc, elementCount2BytesFunc, arrayHeaderClass >::create(), getNumberBases(), and MemoryMapArray< elementT, indexT, cookieVal, versionVal, accessorFunc, setterFunc, elementCount2BytesFunc, arrayHeaderClass >::open().
01255 { 01256 // 01257 // the goal in this section of code is to allow the user 01258 // to either specify a valid binary version of the SNP file, 01259 // or the original text file that it gets created from. 01260 // 01261 // To do this, we basically open, sniff the error message, 01262 // and if it claims it is not a binary version of the file, 01263 // we go ahead and treat it as the text file and use the 01264 // GenomeSequence::populateDBSNP method to load it. 01265 // 01266 // Further checking is really needed to ensure users don't 01267 // mix a dbSNP file for a different reference, since it is really 01268 // easy to do. 01269 // 01270 if (strlen(inputFileName)!=0) 01271 { 01272 std::cerr << "Load dbSNP file '" << inputFileName << "': " << std::flush; 01273 01274 if (dbSNP.open(inputFileName, O_RDONLY)) 01275 { 01276 // 01277 // failed to open, possibly due to bad magic. 01278 // 01279 // this is really awful ... need to have a return 01280 // code that is smart enough to avoid this ugliness: 01281 // 01282 if (dbSNP.getErrorString().find("wrong type of file")==std::string::npos) 01283 { 01284 std::cerr << "Error: " << dbSNP.getErrorString() << std::endl; 01285 exit(1); 01286 } 01287 // 01288 // we have a file, assume we can load it as a text file 01289 // 01290 std::ifstream inputFile; 01291 inputFile.open(inputFileName); 01292 if (inputFile.fail()) 01293 { 01294 std::cerr << "Error: failed to open " << inputFileName << std::endl; 01295 exit(1); 01296 } 01297 01298 std::cerr << "(as text file) "; 01299 01300 // anonymously (RAM resident only) create: 01301 dbSNP.create(getNumberBases()); 01302 01303 // now load it into RAM 01304 populateDBSNP(dbSNP, inputFile); 01305 inputFile.close(); 01306 01307 } 01308 else 01309 { 01310 std::cerr << "(as binary mapped file) "; 01311 } 01312 01313 std::cerr << "DONE!" << std::endl; 01314 return false; 01315 } 01316 else 01317 { 01318 return true; 01319 } 01320 }
| bool GenomeSequence::open | ( | const char * | filename, | |
| int | flags = O_RDONLY | |||
| ) | [inline, virtual] |
open the given file as the genome (no filename munging occurs).
- Parameters:
-
filename the name of the file to open flags pass through to the open() call (O_RDWR lets you modify the contents)
- Returns:
- false for success, true otherwise
Reimplemented from MemoryMapArray< elementT, indexT, cookieVal, versionVal, accessorFunc, setterFunc, elementCount2BytesFunc, arrayHeaderClass >.
Definition at line 315 of file GenomeSequence.h.
References open().
00316 { 00317 _umfaFilename = filename; 00318 return genomeSequenceArray::open(filename, flags); 00319 }
| bool GenomeSequence::open | ( | bool | isColorSpace = false, |
|
| int | flags = O_RDONLY | |||
| ) |
open the reference specified using GenomeSequence::setReferenceName
- Parameters:
-
isColorSpace open the color space reference flags pass through to the open() call (O_RDWR lets you modify the contents)
- Returns:
- false for success, true otherwise
Definition at line 249 of file GenomeSequence.cpp.
Referenced by open().
00250 { 00251 bool rc; 00252 00253 if (isColorSpace) 00254 { 00255 _umfaFilename = _baseFilename + "-cs.umfa"; 00256 } 00257 else 00258 { 00259 _umfaFilename = _baseFilename + "-bs.umfa"; 00260 } 00261 00262 rc = genomeSequenceArray::open(_umfaFilename.c_str(), flags); 00263 if (rc) 00264 { 00265 std::cerr << "GenomeSequence::open: failed to open file " 00266 << _umfaFilename 00267 << std::endl; 00268 return true; 00269 } 00270 00271 _colorSpace = header->_colorSpace; 00272 00273 return false; 00274 }
| char GenomeSequence::operator[] | ( | genomeIndex_t | index | ) | const [inline] |
Return the bases in base space or color space for within range index, ot.
- Parameters:
-
index the array-like index (0 based).
- Returns:
- ACTGN in base space; 0123N for color space; and 'N' for invalid. For color space, index i represents the transition of base at position (i-1) to base at position i
NB: bounds checking here needs to be deprecated - do not assume it will exist - the call must clip reads so that this routine is never called with a index value larger than the genome.
The reason for this is simply that this routine gets called hundreds of billions of time in one run of karma, which will absolutely kill performance. Every single instruction here matters a great, great deal.
Definition at line 516 of file GenomeSequence.h.
References getNumberBases(), and isColorSpace().
00517 { 00518 uint8_t val; 00519 if (index < getNumberBases()) 00520 { 00521 if ((index&1)==0) 00522 { 00523 val = ((uint8_t *) data)[index>>1] & 0xf; 00524 } 00525 else 00526 { 00527 val = (((uint8_t *) data)[index>>1] & 0xf0) >> 4; 00528 } 00529 } 00530 else 00531 { 00532 val = baseNIndex; 00533 } 00534 val = isColorSpace() ? int2colorSpace[val] : int2base[val]; 00535 return val; 00536 }
| void GenomeSequence::setApplication | ( | std::string | application | ) | [inline] |
set the application name in the binary file header
- Parameters:
-
application name of the application
Definition at line 349 of file GenomeSequence.h.
| bool GenomeSequence::setReferenceName | ( | std::string | referenceFilename | ) |
set the reference name that will be used in open()
- Parameters:
-
referenceFilename the name of the reference fasta file to open
- Returns:
- false for success, true otherwise
- See also:
- open()
Definition at line 314 of file GenomeSequence.cpp.
00315 { 00316 00317 if (HAS_SUFFIX(referenceFilename, ".fa")) 00318 { 00319 _referenceFilename = referenceFilename; 00320 _baseFilename = _referenceFilename.substr(0, referenceFilename.size() - 3); 00321 } 00322 else if (HAS_SUFFIX(referenceFilename, ".umfa")) 00323 { 00324 _baseFilename = referenceFilename.substr(0, referenceFilename.size() - 5); 00325 } 00326 else if (HAS_SUFFIX(referenceFilename, "-cs.umfa")) 00327 { 00328 _baseFilename = referenceFilename.substr(0, referenceFilename.size() - 8); 00329 } 00330 else if (HAS_SUFFIX(referenceFilename, "-bs.umfa")) 00331 { 00332 _baseFilename = referenceFilename.substr(0, referenceFilename.size() - 8); 00333 } 00334 else 00335 { 00336 _baseFilename = referenceFilename; 00337 } 00338 _fastaFilename = _baseFilename + ".fa"; 00339 00340 return false; 00341 }
Member Data Documentation
unsigned char GenomeSequence::base2complement [static] |
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"0123NNNNNNNNNNNN"
"NTNGNNNCNNNNNNNN"
"NNNNANNNNNNNNNNN"
"NTNGNNNCNNNNNNNN"
"NNNNANNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
"NNNNNNNNNNNNNNNN"
Definition at line 258 of file GenomeSequence.h.
unsigned char GenomeSequence::base2int [static] |
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\004\005"
"\000\001\002\003\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\000\005\001\005\005\005\002\005\005\005\005\005\005\004\005"
"\005\005\005\005\003\005\005\005\005\005\005\005\005\005\005\005"
"\005\000\005\001\005\005\005\002\005\005\005\005\005\005\004\005"
"\005\005\005\005\003\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
"\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005\005"
Definition at line 253 of file GenomeSequence.h.
The documentation for this class was generated from the following files:
- general/GenomeSequence.h
- general/GenomeSequence.cpp
 1.6.3
1.6.3