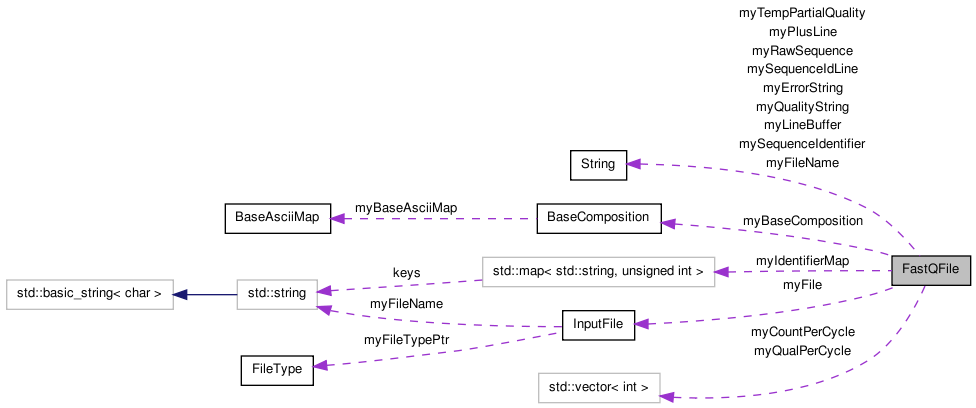
FastQFile Class Reference
Class for reading/validating a fastq file. More...
#include <FastQFile.h>
Public Member Functions | |
| FastQFile (int minReadLength=10, int numPrintableErrors=20) | |
| Constructor. | |
| void | disableMessages () |
| Disable messages - do not write to cout. | |
| void | enableMessages () |
| Enable messages - write to cout. | |
| void | disableSeqIDCheck () |
| Disable Unique Sequence ID checking (Unique Sequence ID checking is enabled by default). | |
| void | enableSeqIDCheck () |
| Enable Unique Sequence ID checking. | |
| void | setMaxErrors (int maxErrors) |
| Set the number of errors after which to quit reading/validating a file, defaults to -1. | |
| FastQStatus::Status | openFile (const char *fileName, BaseAsciiMap::SPACE_TYPE spaceType=BaseAsciiMap::UNKNOWN) |
| Open a FastQFile. | |
| FastQStatus::Status | closeFile () |
| Close a FastQFile. | |
| bool | isOpen () |
| Check to see if the file is open. | |
| bool | isEof () |
| Check to see if the file is at the end of the file. | |
| bool | keepReadingFile () |
| Returns whether or not to keep reading the file, it stops reading (false) if eof or there is a problem reading the file. | |
| FastQStatus::Status | validateFastQFile (const String &filename, bool printBaseComp, BaseAsciiMap::SPACE_TYPE spaceType, bool printQualAvg=false) |
| Validate the specified fastq file. | |
| FastQStatus::Status | readFastQSequence () |
| Read 1 FastQSequence, validating it. | |
| BaseAsciiMap::SPACE_TYPE | getSpaceType () |
| Get the space type used for this file. | |
Public Attributes | |
Public Sequence Line variables. | |
| String | myRawSequence |
| String | mySequenceIdLine |
| String | mySequenceIdentifier |
| String | myPlusLine |
| String | myQualityString |
Detailed Description
Class for reading/validating a fastq file.
Definition at line 29 of file FastQFile.h.
Constructor & Destructor Documentation
| FastQFile::FastQFile | ( | int | minReadLength = 10, |
|
| int | numPrintableErrors = 20 | |||
| ) |
Constructor.
/param minReadLength The minimum length that a base sequence must be for it to be valid.
- Parameters:
-
numPrintableErrors The maximum number of errors that should be reported in detail before suppressing the errors.
Definition at line 30 of file FastQFile.cpp.
00031 : myFile(NULL), 00032 myBaseComposition(), 00033 myQualPerCycle(), 00034 myCountPerCycle(), 00035 myCheckSeqID(true), 00036 myMinReadLength(minReadLength), 00037 myNumPrintableErrors(numPrintableErrors), 00038 myMaxErrors(-1), 00039 myDisableMessages(false), 00040 myFileProblem(false) 00041 { 00042 // Reset the member data. 00043 reset(); 00044 }
Member Function Documentation
| void FastQFile::disableSeqIDCheck | ( | ) |
Disable Unique Sequence ID checking (Unique Sequence ID checking is enabled by default).
Definition at line 61 of file FastQFile.cpp.
| void FastQFile::enableSeqIDCheck | ( | ) |
Enable Unique Sequence ID checking.
(Unique Sequence ID checking is enabled by default).
Definition at line 69 of file FastQFile.cpp.
| bool FastQFile::keepReadingFile | ( | ) |
Returns whether or not to keep reading the file, it stops reading (false) if eof or there is a problem reading the file.
Definition at line 184 of file FastQFile.cpp.
References isEof().
Referenced by validateFastQFile().
00185 { 00186 if(isEof() || myFileProblem) 00187 { 00188 return(false); 00189 } 00190 return(true); 00191 }
| FastQStatus::Status FastQFile::openFile | ( | const char * | fileName, | |
| BaseAsciiMap::SPACE_TYPE | spaceType = BaseAsciiMap::UNKNOWN | |||
| ) |
Open a FastQFile.
Use the specified SPACE_TYPE to determine BASE, COLOR, or UNKNOWN.
Definition at line 83 of file FastQFile.cpp.
References closeFile(), FastQStatus::FASTQ_OPEN_ERROR, FastQStatus::FASTQ_SUCCESS, ifopen(), BaseComposition::resetBaseMapType(), and BaseComposition::setBaseMapType().
Referenced by validateFastQFile().
00085 { 00086 // reset the member data. 00087 reset(); 00088 00089 myBaseComposition.resetBaseMapType(); 00090 myBaseComposition.setBaseMapType(spaceType); 00091 myQualPerCycle.clear(); 00092 myCountPerCycle.clear(); 00093 00094 FastQStatus::Status status = FastQStatus::FASTQ_SUCCESS; 00095 00096 // Close the file if there is already one open - checked by close. 00097 status = closeFile(); 00098 if(status == FastQStatus::FASTQ_SUCCESS) 00099 { 00100 // Successfully closed a previously opened file if there was one. 00101 00102 // Open the file 00103 myFile = ifopen(fileName, "rt"); 00104 myFileName = fileName; 00105 00106 if(myFile == NULL) 00107 { 00108 // Failed to open the file. 00109 status = FastQStatus::FASTQ_OPEN_ERROR; 00110 } 00111 } 00112 00113 if(status != FastQStatus::FASTQ_SUCCESS) 00114 { 00115 // Failed to open the file. 00116 std::string errorMessage = "ERROR: Failed to open file: "; 00117 errorMessage += fileName; 00118 logMessage(errorMessage.c_str()); 00119 } 00120 return(status); 00121 }
| void FastQFile::setMaxErrors | ( | int | maxErrors | ) |
Set the number of errors after which to quit reading/validating a file, defaults to -1.
- Parameters:
-
maxErrors # of errors before quitting, -1 indicates to not quit until the entire file has been read/validated (default), 0 indicates to quit without reading/validating anything.
Definition at line 76 of file FastQFile.cpp.
| FastQStatus::Status FastQFile::validateFastQFile | ( | const String & | filename, | |
| bool | printBaseComp, | |||
| BaseAsciiMap::SPACE_TYPE | spaceType, | |||
| bool | printQualAvg = false | |||
| ) |
Validate the specified fastq file.
- Parameters:
-
filename fastq file to be validated. printBaseComp whether or not to print the base composition for the file. true means print it, false means do not. spaceType the spaceType to use for validation - BASE_SPACE, COLOR_SPACE, or UNKNOWN (UNKNOWN means to determine the spaceType to validate against from the first character of the first sequence). printQualAvg whether or not to print the quality averages for the file. true means to print it, false (default) means do not.
- Returns:
- the fastq validation status, SUCCESS on a successfully validated fastq file.
Definition at line 195 of file FastQFile.cpp.
References closeFile(), FastQStatus::FASTQ_INVALID, FastQStatus::FASTQ_NO_SEQUENCE_ERROR, FastQStatus::FASTQ_OPEN_ERROR, FastQStatus::FASTQ_SUCCESS, keepReadingFile(), openFile(), BaseComposition::print(), and readFastQSequence().
00199 { 00200 // Open the fastqfile. 00201 if(openFile(filename, spaceType) != FastQStatus::FASTQ_SUCCESS) 00202 { 00203 // Failed to open the specified file. 00204 return(FastQStatus::FASTQ_OPEN_ERROR); 00205 } 00206 00207 // Track the total number of sequences that were validated. 00208 int numSequences = 0; 00209 00210 // Keep reading the file until there are no more fastq sequences to process 00211 // and not configured to quit after a certain number of errors or there 00212 // has not yet been that many errors. 00213 // Or exit if there is a problem reading the file. 00214 FastQStatus::Status status = FastQStatus::FASTQ_SUCCESS; 00215 while (keepReadingFile() && 00216 ((myMaxErrors == -1) || (myMaxErrors > myNumErrors))) 00217 { 00218 // Validate one sequence. This call will read all the lines for 00219 // one sequence. 00220 status = readFastQSequence(); 00221 if((status == FastQStatus::FASTQ_SUCCESS) || (status == FastQStatus::FASTQ_INVALID)) 00222 { 00223 // Read a sequence and it is either valid or invalid, but 00224 // either way, a sequence was read, so increment the sequence count. 00225 ++numSequences; 00226 } 00227 else 00228 { 00229 // Other error, so break out of processing. 00230 break; 00231 } 00232 } 00233 00234 // Report Base Composition Statistics. 00235 if(printBaseComp) 00236 { 00237 myBaseComposition.print(); 00238 } 00239 00240 if(printQualAvg) 00241 { 00242 printAvgQual(); 00243 } 00244 00245 std::string finishMessage = "Finished processing "; 00246 finishMessage += myFileName.c_str(); 00247 char buffer[100]; 00248 if(sprintf(buffer, 00249 " with %u lines containing %d sequences.", 00250 myLineNum, numSequences) > 0) 00251 { 00252 finishMessage += buffer; 00253 logMessage(finishMessage.c_str()); 00254 } 00255 if(sprintf(buffer, 00256 "There were a total of %d errors.", 00257 myNumErrors) > 0) 00258 { 00259 logMessage(buffer); 00260 } 00261 00262 // Close the input file. 00263 FastQStatus::Status closeStatus = closeFile(); 00264 00265 if((status != FastQStatus::FASTQ_SUCCESS) && (status != FastQStatus::FASTQ_INVALID)) 00266 { 00267 // Stopped validating due to some error other than invalid, so 00268 // return that error. 00269 return(status); 00270 } 00271 else if(myNumErrors == 0) 00272 { 00273 // No errors, check to see if there were any sequences. 00274 // Finished processing all of the sequences in the file. 00275 // If there are no sequences, report an error. 00276 if(numSequences == 0) 00277 { 00278 // Empty file, return error. 00279 logMessage("ERROR: No FastQSequences in the file."); 00280 return(FastQStatus::FASTQ_NO_SEQUENCE_ERROR); 00281 } 00282 return(FastQStatus::FASTQ_SUCCESS); 00283 } 00284 else 00285 { 00286 // The file is invalid. But check the close status. If the close 00287 // failed, it means there is a problem with the file itself not just 00288 // with validation, so the close failure should be returned. 00289 if(closeStatus != FastQStatus::FASTQ_SUCCESS) 00290 { 00291 return(closeStatus); 00292 } 00293 return(FastQStatus::FASTQ_INVALID); 00294 } 00295 }
The documentation for this class was generated from the following files:
- fastq/FastQFile.h
- fastq/FastQFile.cpp
 1.6.3
1.6.3