GenomeSequence Class Reference
Create/Access/Modify/Load Genome Sequences stored as binary mapped files. More...
#include <GenomeSequence.h>
Public Member Functions | |
| GenomeSequence () | |
| Simple constructor - no implicit file open. | |
| void | constructorClear () |
| GenomeSequence (std::string &referenceFilename) | |
| attempt to open an existing sequence | |
| GenomeSequence (const char *referenceFilename) | |
| Smarter constructor - attempt to open an existing sequence. | |
| ~GenomeSequence () | |
| Close the file if open and destroy the object. | |
| bool | open (bool isColorSpace=false, int flags=O_RDONLY) |
| open the reference specified using GenomeSequence::setReferenceName | |
| bool | open (const char *filename, int flags=O_RDONLY) |
| open the given file as the genome (no filename munging occurs). | |
| bool | create (bool isColor=false) |
| void | setProgressStream (std::ostream &progressStream) |
| if set, then show progress when creating and pre-fetching | |
| void | setColorSpace (bool colorSpace) |
| void | setSearchCommonFileSuffix (bool searchCommonFileSuffix) |
| void | setCreateOverwrite (bool createOverwrite) |
| bool | loadFastaData (const char *filename) |
| bool | setReferenceName (std::string referenceFilename) |
| set the reference name that will be used in open() | |
| void | setApplication (std::string application) |
| set the application name in the binary file header | |
| const std::string & | getFastaName () const |
| const std::string & | getReferenceName () const |
| bool | isColorSpace () const |
| tell us if we are a color space reference or not | |
| genomeIndex_t | getNumberBases () const |
| return the number of bases represented in this reference | |
| int | getChromosome (genomeIndex_t position) const |
| given a whole genome index, get the chromosome it is located in | |
| int | getChromosome (const char *chromosomeName) const |
| given a chromosome name, return the chromosome index | |
| int | getChromosomeCount () const |
| Return the number of chromosomes in the genome. | |
| genomeIndex_t | getChromosomeStart (int chromosomeIndex) const |
| given a chromosome, return the genome base it starts in | |
| genomeIndex_t | getChromosomeSize (int chromosomeIndex) const |
| given a chromosome, return its size in bases | |
| genomeIndex_t | getGenomePosition (const char *chromosomeName, unsigned int chromosomeIndex) const |
| given a chromosome name and position, return the genome position | |
| genomeIndex_t | getGenomePosition (int chromosome, unsigned int chromosomeIndex) const |
| given a chromosome index and position, return the genome position | |
| genomeIndex_t | getGenomePosition (const char *chromosomeName) const |
| given the chromosome name, get the corresponding 0 based genome index for the start of that chromosome | |
| genomeIndex_t | getGenomePosition (int chromosomeIndex) const |
| const std::string & | getBaseFilename () const |
| const char * | getChromosomeName (int chromosomeIndex) const |
| void | setDebugFlag (bool d) |
| genomeIndex_t | sequenceLength () const |
| const char * | chromosomeName (int chr) const |
| void | sanityCheck (MemoryMap &fasta) const |
| std::string | IntegerToSeq (unsigned int n, unsigned int wordsize) const |
| bool | wordMatch (unsigned int index, std::string &word) const |
| bool | printNearbyWords (unsigned int index, unsigned int variance, std::string &word) const |
| char | BasePair (char c) const |
| void | dumpSequenceSAMDictionary (std::ostream &) const |
| void | dumpHeaderTSV (std::ostream &) const |
| char | operator[] (genomeIndex_t index) const |
| Return the bases in base space or color space for within range index, ot. | |
| char | getBase (const char *chromosomeName, unsigned int chromosomeIndex) const |
| given a chromosome name and 1-based position, return the reference base. | |
| uint8_t | getInteger (genomeIndex_t index) const |
| void | set (genomeIndex_t index, char value) |
| uint8_t * | getDataPtr (genomeIndex_t index) |
| obtain the pointer to the raw data for other access methods | |
| void | getReverseRead (std::string &read) |
| void | getReverseRead (String &read) |
| int | debugPrintReadValidation (std::string &read, std::string &quality, char direction, genomeIndex_t readLocation, int sumQuality, int mismatchCount, bool recurse=true) |
| void | getString (std::string &str, int chromosome, uint32_t index, int baseCount) const |
| void | getString (String &str, int chromosome, uint32_t index, int baseCount) const |
| void | getString (std::string &str, genomeIndex_t index, int baseCount) const |
| void | getString (String &str, genomeIndex_t index, int baseCount) const |
| void | getHighLightedString (std::string &str, genomeIndex_t index, int baseCount, genomeIndex_t highLightStart, genomeIndex_t highLightEnd) const |
| void | print30 (genomeIndex_t) const |
| genomeIndex_t | simpleLocalAligner (std::string &read, std::string &quality, genomeIndex_t index, int windowSize) const |
| int | getMismatchCount (std::string &read, genomeIndex_t location, char exclude='\0') const |
| Return the mismatch count, disregarding CIGAR strings. | |
| int | getSumQ (std::string &read, std::string &qualities, genomeIndex_t location) const |
| brute force sumQ - no sanity checking | |
| void | getMismatchHatString (std::string &result, const std::string &read, genomeIndex_t location) const |
| void | getMismatchString (std::string &result, const std::string &read, genomeIndex_t location) const |
| void | getChromosomeAndIndex (std::string &, genomeIndex_t) const |
| void | getChromosomeAndIndex (String &, genomeIndex_t) const |
| bool | checkRead (std::string &read, std::string &qualities, std::string &cigar, int &sumQ, int &gapOpenCount, int &gapExtendCount, int &gapDeleteCount, std::string &result) const |
| check a SAM format read, using phred quality scores and the CIGAR string to determine if it is correct. | |
| bool | populateDBSNP (mmapArrayBool_t &dbSNP, IFILE inputFile) const |
| bool | loadDBSNP (mmapArrayBool_t &dbSNP, const char *inputFileName) const |
| user friendly dbSNP loader. | |
Detailed Description
Create/Access/Modify/Load Genome Sequences stored as binary mapped files.
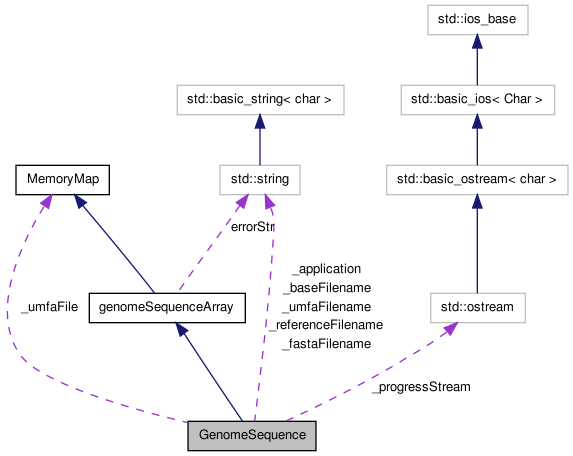
GenomeSequence is designed to be a high performance shared access reference object.
It is implemented as a MemoryMapArray template object with unsigned 8 bit ints, each of which stores two bases. Although 2 bits could be used, most references have more than four symbols (usually at least including 'N', indicating an unknown or masked out base).
Normal use of this class follows these steps:
- create the reference
- instantiate the GenomeSequence class object
- create the actual file (memory mapped) that is to hold the data
- populate the data using GenomeSequence::set
- use the reference
- use the reference by instantiating a GenomeSequence object
- either use the constructor with the reference filename
- or use GenomeSequence::setReferenceName() followed by open
- access the bases via the overloaded array operator []
- check sequence length by using GenomeSequence::getNumberBases()
- accessing chromosomes in the reference
- you typically will need to know about the chromosomes in the sequence
- see methods and docs with prefix 'getChromosome'
Sharing is accomplished using the mmap() function via the MemoryMap base class. This allows a potentially large genome reference to be shared among a number of simultaneously executing instances of one or more programs sharing the same reference.
Definition at line 99 of file GenomeSequence.h.
Constructor & Destructor Documentation
| GenomeSequence::GenomeSequence | ( | std::string & | referenceFilename | ) | [inline] |
attempt to open an existing sequence
- Parameters:
-
referenceFilename the name of the reference fasta file to open debug if true, additional debug information is printed
Definition at line 128 of file GenomeSequence.h.
| GenomeSequence::GenomeSequence | ( | const char * | referenceFilename | ) | [inline] |
Smarter constructor - attempt to open an existing sequence.
- Parameters:
-
referenceFilename the name of the reference fasta file to open debug if true, additional debug information is printed
Definition at line 138 of file GenomeSequence.h.
Member Function Documentation
| bool GenomeSequence::checkRead | ( | std::string & | read, | |
| std::string & | qualities, | |||
| std::string & | cigar, | |||
| int & | sumQ, | |||
| int & | gapOpenCount, | |||
| int & | gapExtendCount, | |||
| int & | gapDeleteCount, | |||
| std::string & | result | |||
| ) | const |
check a SAM format read, using phred quality scores and the CIGAR string to determine if it is correct.
- Parameters:
-
read the read in base space qualities the phred encoded qualities (Sanger, not Illumina) cigar the SAM file CIGAR column sumQ if >0 on entry, is checked against the computed sumQ insertions count of insertions found in
| char GenomeSequence::getBase | ( | const char * | chromosomeName, | |
| unsigned int | chromosomeIndex | |||
| ) | const [inline] |
given a chromosome name and 1-based position, return the reference base.
- Parameters:
-
chromosomeName name of the chromosome - exact match only chromosomeIndex 1-based chromosome position
- Returns:
- reference base at the above chromosome position
Definition at line 396 of file GenomeSequence.h.
References getGenomePosition().
Referenced by PileupElement::getRefBase().
00398 { 00399 genomeIndex_t index = 00400 getGenomePosition(chromosomeName, chromosomeIndex); 00401 if(index == INVALID_GENOME_INDEX) 00402 { 00403 // Invalid position, so return 'N' 00404 return('N'); 00405 } 00406 return((*this)[index]); 00407 }
| int GenomeSequence::getChromosome | ( | const char * | chromosomeName | ) | const |
given a chromosome name, return the chromosome index
This is done via a linear search of the chromosome table in the header of the mapped file, so it is O(N)
- Parameters:
-
chromosomeName the name of the chromosome - exact match only
- Returns:
- 0-based index into chromosome table - INVALID_CHROMOSOME_INDEX if error
Definition at line 877 of file GenomeSequence.cpp.
| int GenomeSequence::getChromosome | ( | genomeIndex_t | position | ) | const |
given a whole genome index, get the chromosome it is located in
This is done via a binary search of the chromosome table in the header of the mapped file, so it is O(log(N))
- Parameters:
-
0-based position the base in the genome
- Returns:
- 0-based index into chromosome table - INVALID_CHROMOSOME_INDEX if error
Definition at line 800 of file GenomeSequence.cpp.
Referenced by getGenomePosition().
00801 { 00802 if (position == INVALID_GENOME_INDEX) return INVALID_CHROMOSOME_INDEX; 00803 00804 if (header->_chromosomeCount == 0) 00805 return INVALID_CHROMOSOME_INDEX; 00806 00807 int start = 0; 00808 int stop = header->_chromosomeCount - 1; 00809 00810 // eliminate case where position is in the last chromosome, since the loop 00811 // below falls off the end of the list if it in the last one. 00812 00813 if (position > header->_chromosomes[stop].start) 00814 return (stop); 00815 00816 while (start <= stop) 00817 { 00818 int middle = (start + stop) / 2; 00819 00820 if (position >= header->_chromosomes[middle].start && position < header->_chromosomes[middle + 1].start) 00821 return middle; 00822 00823 if (position == header->_chromosomes[middle + 1].start) 00824 return (middle + 1); 00825 00826 if (position > header->_chromosomes[middle + 1].start) 00827 start = middle + 1; 00828 00829 if (position < header->_chromosomes[middle].start) 00830 stop = middle - 1; 00831 } 00832 00833 return -1; 00834 }
| int GenomeSequence::getChromosomeCount | ( | ) | const |
Return the number of chromosomes in the genome.
- Returns:
- number of chromosomes in the genome
Definition at line 794 of file GenomeSequence.cpp.
| genomeIndex_t GenomeSequence::getChromosomeSize | ( | int | chromosomeIndex | ) | const [inline] |
given a chromosome, return its size in bases
- Parameters:
-
0-based chromosome index
- Returns:
- size of the chromosome in bases
Definition at line 264 of file GenomeSequence.h.
| genomeIndex_t GenomeSequence::getChromosomeStart | ( | int | chromosomeIndex | ) | const [inline] |
given a chromosome, return the genome base it starts in
- Parameters:
-
0-based chromosome index
- Returns:
- 0-based genome index of the base that starts the chromosome
Definition at line 254 of file GenomeSequence.h.
| uint8_t* GenomeSequence::getDataPtr | ( | genomeIndex_t | index | ) | [inline] |
obtain the pointer to the raw data for other access methods
this is a fairly ugly hack to reach into the raw genome vector, get the byte that encodes two bases, and return it. This is used by karma ReadIndexer::getSumQ to compare genome matchines by byte (two bases at a time) to speed it up.
Definition at line 430 of file GenomeSequence.h.
| genomeIndex_t GenomeSequence::getGenomePosition | ( | int | chromosome, | |
| unsigned int | chromosomeIndex | |||
| ) | const |
given a chromosome index and position, return the genome position
- Parameters:
-
chromosome index of the chromosome chromosomeIndex 1-based chromosome position
- Returns:
- genome index of the above chromosome position
Definition at line 851 of file GenomeSequence.cpp.
| genomeIndex_t GenomeSequence::getGenomePosition | ( | const char * | chromosomeName, | |
| unsigned int | chromosomeIndex | |||
| ) | const |
given a chromosome name and position, return the genome position
- Parameters:
-
chromosomeName name of the chromosome - exact match only chromosomeIndex 1-based chromosome position
- Returns:
- genome index of the above chromosome position
Definition at line 842 of file GenomeSequence.cpp.
Referenced by SamTags::createMDTag(), getBase(), SamQuerySeqWithRefIter::reset(), SamQuerySeqWithRef::seqWithEquals(), and SamQuerySeqWithRef::seqWithoutEquals().
00845 { 00846 genomeIndex_t i = getGenomePosition(chromosomeName); 00847 if (i == INVALID_GENOME_INDEX) return INVALID_GENOME_INDEX; 00848 return i + chromosomeIndex - 1; 00849 }
| int GenomeSequence::getMismatchCount | ( | std::string & | read, | |
| genomeIndex_t | location, | |||
| char | exclude = '\0' | |||
| ) | const [inline] |
Return the mismatch count, disregarding CIGAR strings.
- Parameters:
-
read is the sequence we're counting mismatches in location is where in the genmoe we start comparing exclude is a wildcard character (e.g. '.' or 'N')
- Returns:
- number of bases that don't match the reference, except those that match exclude
Definition at line 496 of file GenomeSequence.h.
| genomeIndex_t GenomeSequence::getNumberBases | ( | ) | const [inline] |
return the number of bases represented in this reference
- Returns:
- count of bases
Definition at line 224 of file GenomeSequence.h.
Referenced by loadDBSNP(), and operator[]().
| int GenomeSequence::getSumQ | ( | std::string & | read, | |
| std::string & | qualities, | |||
| genomeIndex_t | location | |||
| ) | const [inline] |
brute force sumQ - no sanity checking
- Parameters:
-
read shotgun sequencer read string qualities phred quality string of same length location the alignment location to check sumQ
Definition at line 509 of file GenomeSequence.h.
| bool GenomeSequence::isColorSpace | ( | ) | const [inline] |
tell us if we are a color space reference or not
- Returns:
- true if colorspace, false otherwise
Definition at line 217 of file GenomeSequence.h.
Referenced by operator[]().
| bool GenomeSequence::loadDBSNP | ( | mmapArrayBool_t & | dbSNP, | |
| const char * | inputFileName | |||
| ) | const |
user friendly dbSNP loader.
- Parameters:
-
inputFileName may be empty, point to a text file or a dbSNP vector file
In all cases, dbSNP is returned the same length as this genome.
When no SNPs are loaded, all values are false.
When a text file is given, the file is parsed with two space separated columns - the first column is the chromosome name, and the second is the 1-based chromosome position of the SNP.
- Returns:
- false if a dbSNP file was correctly loaded, true otherwise
Definition at line 1364 of file GenomeSequence.cpp.
References MemoryMapArray< elementT, indexT, cookieVal, versionVal, accessorFunc, setterFunc, elementCount2BytesFunc, arrayHeaderClass >::create(), getNumberBases(), ifclose(), ifopen(), and MemoryMapArray< elementT, indexT, cookieVal, versionVal, accessorFunc, setterFunc, elementCount2BytesFunc, arrayHeaderClass >::open().
01367 { 01368 // 01369 // the goal in this section of code is to allow the user 01370 // to either specify a valid binary version of the SNP file, 01371 // or the original text file that it gets created from. 01372 // 01373 // To do this, we basically open, sniff the error message, 01374 // and if it claims it is not a binary version of the file, 01375 // we go ahead and treat it as the text file and use the 01376 // GenomeSequence::populateDBSNP method to load it. 01377 // 01378 // Further checking is really needed to ensure users don't 01379 // mix a dbSNP file for a different reference, since it is really 01380 // easy to do. 01381 // 01382 if (strlen(inputFileName)!=0) 01383 { 01384 std::cerr << "Load dbSNP file '" << inputFileName << "': " << std::flush; 01385 01386 if (dbSNP.open(inputFileName, O_RDONLY)) 01387 { 01388 // 01389 // failed to open, possibly due to bad magic. 01390 // 01391 // this is really awful ... need to have a return 01392 // code that is smart enough to avoid this ugliness: 01393 // 01394 if (dbSNP.getErrorString().find("wrong type of file")==std::string::npos) 01395 { 01396 std::cerr << "Error: " << dbSNP.getErrorString() << std::endl; 01397 exit(1); 01398 } 01399 // 01400 // we have a file, assume we can load it as a text file 01401 // 01402 IFILE inputFile = ifopen(inputFileName, "r"); 01403 if(inputFile == NULL) 01404 { 01405 std::cerr << "Error: failed to open " << inputFileName << std::endl; 01406 exit(1); 01407 } 01408 01409 std::cerr << "(as text file) "; 01410 01411 // anonymously (RAM resident only) create: 01412 dbSNP.create(getNumberBases()); 01413 01414 // now load it into RAM 01415 populateDBSNP(dbSNP, inputFile); 01416 ifclose(inputFile); 01417 01418 } 01419 else 01420 { 01421 std::cerr << "(as binary mapped file) "; 01422 } 01423 01424 std::cerr << "DONE!" << std::endl; 01425 return false; 01426 } 01427 else 01428 { 01429 return true; 01430 } 01431 }
| bool GenomeSequence::open | ( | const char * | filename, | |
| int | flags = O_RDONLY | |||
| ) | [inline, virtual] |
open the given file as the genome (no filename munging occurs).
- Parameters:
-
filename the name of the file to open flags pass through to the open() call (O_RDWR lets you modify the contents)
- Returns:
- false for success, true otherwise
Reimplemented from MemoryMapArray< elementT, indexT, cookieVal, versionVal, accessorFunc, setterFunc, elementCount2BytesFunc, arrayHeaderClass >.
Definition at line 159 of file GenomeSequence.h.
00160 { 00161 _umfaFilename = filename; 00162 #if 0 00163 if(genomeSequenceArray::open(filename, flags)) { 00164 // 00165 // if load of the umfa (binary encoded file) fails, attempt 00166 // to load it as a fasta file. 00167 // 00168 return loadFasta(); 00169 } 00170 #endif 00171 return false; 00172 }
| bool GenomeSequence::open | ( | bool | isColorSpace = false, |
|
| int | flags = O_RDONLY | |||
| ) |
open the reference specified using GenomeSequence::setReferenceName
- Parameters:
-
isColorSpace open the color space reference flags pass through to the open() call (O_RDWR lets you modify the contents)
- Returns:
- false for success, true otherwise
Definition at line 178 of file GenomeSequence.cpp.
00179 { 00180 bool rc; 00181 00182 if (isColorSpace) 00183 { 00184 _umfaFilename = _baseFilename + "-cs.umfa"; 00185 } 00186 else 00187 { 00188 _umfaFilename = _baseFilename + "-bs.umfa"; 00189 } 00190 00191 if(access(_umfaFilename.c_str(), R_OK) != 0) 00192 { 00193 // umfa file doesn't exist, so try to create it. 00194 if(create(isColorSpace)) 00195 { 00196 // Couldon't access or create the umfa. 00197 std::cerr << "GenomeSequence::open: failed to open file " 00198 << _umfaFilename 00199 << " also failed creating it." 00200 << std::endl; 00201 return true; 00202 } 00203 } 00204 00205 rc = genomeSequenceArray::open(_umfaFilename.c_str(), flags); 00206 if (rc) 00207 { 00208 std::cerr << "GenomeSequence::open: failed to open file " 00209 << _umfaFilename 00210 << std::endl; 00211 return true; 00212 } 00213 00214 _colorSpace = header->_colorSpace; 00215 00216 return false; 00217 }
| char GenomeSequence::operator[] | ( | genomeIndex_t | index | ) | const [inline] |
Return the bases in base space or color space for within range index, ot.
- Parameters:
-
index the array-like index (0 based).
- Returns:
- ACTGN in base space; 0123N for color space; and 'N' for invalid. For color space, index i represents the transition of base at position (i-1) to base at position i
NB: bounds checking here needs to be deprecated - do not assume it will exist - the call must clip reads so that this routine is never called with a index value larger than the genome.
The reason for this is simply that this routine gets called hundreds of billions of time in one run of karma, which will absolutely kill performance. Every single instruction here matters a great, great deal.
Definition at line 369 of file GenomeSequence.h.
References BaseAsciiMap::baseNIndex, getNumberBases(), BaseAsciiMap::int2base, BaseAsciiMap::int2colorSpace, and isColorSpace().
00370 { 00371 uint8_t val; 00372 if (index < getNumberBases()) 00373 { 00374 if ((index&1)==0) 00375 { 00376 val = ((uint8_t *) data)[index>>1] & 0xf; 00377 } 00378 else 00379 { 00380 val = (((uint8_t *) data)[index>>1] & 0xf0) >> 4; 00381 } 00382 } 00383 else 00384 { 00385 val = BaseAsciiMap::baseNIndex; 00386 } 00387 val = isColorSpace() ? BaseAsciiMap::int2colorSpace[val] : 00388 BaseAsciiMap::int2base[val]; 00389 return val; 00390 }
| void GenomeSequence::setApplication | ( | std::string | application | ) | [inline] |
set the application name in the binary file header
- Parameters:
-
application name of the application
Definition at line 202 of file GenomeSequence.h.
| bool GenomeSequence::setReferenceName | ( | std::string | referenceFilename | ) |
set the reference name that will be used in open()
- Parameters:
-
referenceFilename the name of the reference fasta file to open
- Returns:
- false for success, true otherwise
- See also:
- open()
Definition at line 260 of file GenomeSequence.cpp.
00261 { 00262 00263 if (HAS_SUFFIX(referenceFilename, ".fa")) 00264 { 00265 _referenceFilename = referenceFilename; 00266 _baseFilename = _referenceFilename.substr(0, referenceFilename.size() - 3); 00267 } 00268 else if (HAS_SUFFIX(referenceFilename, ".umfa")) 00269 { 00270 _baseFilename = referenceFilename.substr(0, referenceFilename.size() - 5); 00271 } 00272 else if (HAS_SUFFIX(referenceFilename, "-cs.umfa")) 00273 { 00274 _baseFilename = referenceFilename.substr(0, referenceFilename.size() - 8); 00275 } 00276 else if (HAS_SUFFIX(referenceFilename, "-bs.umfa")) 00277 { 00278 _baseFilename = referenceFilename.substr(0, referenceFilename.size() - 8); 00279 } 00280 else 00281 { 00282 _baseFilename = referenceFilename; 00283 } 00284 _fastaFilename = _baseFilename + ".fa"; 00285 00286 if (HAS_SUFFIX(referenceFilename, ".fasta")) 00287 { 00288 _referenceFilename = referenceFilename; 00289 _baseFilename = _referenceFilename.substr(0, referenceFilename.size() - 6); 00290 _fastaFilename = _baseFilename + ".fasta"; 00291 } 00292 00293 return false; 00294 }
The documentation for this class was generated from the following files:
- general/GenomeSequence.h
- general/GenomeSequence.cpp
 1.6.3
1.6.3