AMD Gene Consortium Study of Age Related Macular Degeneration
Meta-Analysis Tables
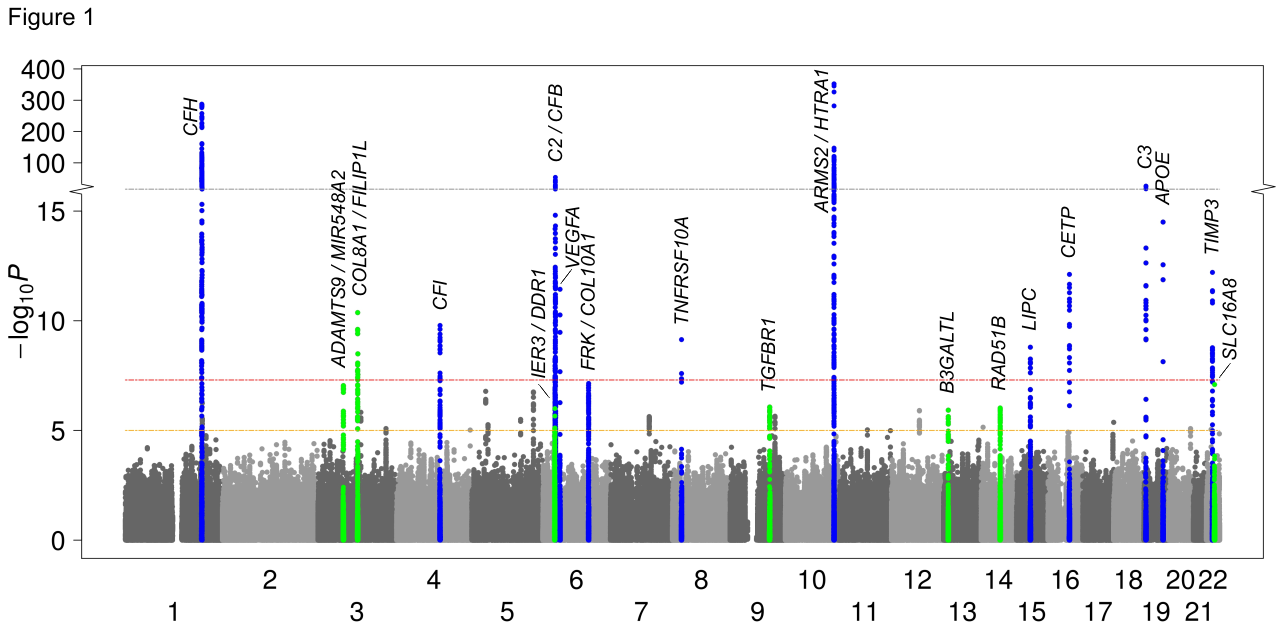
The three tables below summarize the results of the meta-analysis described
in Fritsche et al, Nature Genetics (2013). Each table includes seven
columns: the rs# for each SNP evaluated, the two alleles of the SNP in genome-build
37, the number of cases and controls evaluated for the SNP,
the combined p-value for the SNP, the overall direction of effect
(a positive statistic indicates allele 1 was associated with increased
risk of macular degeneration). The files included both genotyped and imputed SNPs
and are provided in a compressed text file to conserve space.
To browse results in a user friendly fashion, try LocusZoom, and
request the option to plot published GWAS results.
RESULT FILES
Advanced AMD vs. Control Analysis GZIP Archive
Geographic Atrophy vs. Control Analysis GZIP Archive
Neovascular Disease vs. Control Analysis GZIP Archive
Columns in Result Files
| Column Label | Description |
| Marker | This is the marker name. |
| Allele1 | This the allele used as the effect allele. |
| Allele2 | This is the "other" allele. |
| Ncases | The number of cases analysed for this marker. |
| Ncontrols | The number of controls analysed for this marker. |
| GC.Pvalue | Meta-analysis p-value, after genomic control correction. |
| Overall | Overall direction of effect. Positive values indicate allele1 was associated with increased disease risk. |
NOTE:These files were updated on April 24, 2013 to fix a formatting
issue; that resulted in wrong direction of effect for several SNPs.
CITATION:If you use these GWAS results in any published work, please cite the following manuscript:
Fritsche, L.G. et al. Seven new loci associated with age-related macular degeneration. Nat Genet. 2013 Apr;45(4):433-9, 439e1-2. doi: 10.1038/ng.2578. PubMed PMID: 23455636.
Please e-mail Goncalo Abecasis or
Lars Fritsche with comments or
suggestions.
| 


