SamFile Class Reference
Allows the user to easily read/write a SAM/BAM file. More...
#include <SamFile.h>
Public Types | |
| enum | OpenType { READ, WRITE } |
Enum for indicating whether to open the file for read or write. More... | |
| enum | SortedType { UNSORTED = 0, FLAG, COORDINATE, QUERY_NAME } |
Enum for indicating the type of sort expected in the file. More... | |
Public Member Functions | |
| SamFile () | |
| Default Constructor, initializes the variables, but does not open any files. | |
| SamFile (ErrorHandler::HandlingType errorHandlingType) | |
| Constructor that sets the error handling type. | |
| SamFile (const char *filename, OpenType mode) | |
| Constructor that opens the specified file based on the specified mode (READ/WRITE), aborts if the file could not be opened. | |
| SamFile (const char *filename, OpenType mode, ErrorHandler::HandlingType errorHandlingType) | |
| Constructor that opens the specified file based on the specified mode (READ/WRITE) and handles errors per the specified handleType. | |
| SamFile (const char *filename, OpenType mode, SamFileHeader *header) | |
| Constructor that opens the specified file based on the specified mode (READ/WRITE) and reads the header, aborts if the file could not be opened or the header not read. | |
| SamFile (const char *filename, OpenType mode, ErrorHandler::HandlingType errorHandlingType, SamFileHeader *header) | |
| Constructor that opens the specified file based on the specified mode (READ/WRITE) and reads the header, handling errors per the specified handleType. | |
| virtual | ~SamFile () |
| Destructor. | |
| bool | OpenForRead (const char *filename, SamFileHeader *header=NULL) |
| Open a sam/bam file for reading with the specified filename, determing the type of file and SAM/BAM by reading the file (if not stdin). | |
| bool | OpenForWrite (const char *filename, SamFileHeader *header=NULL) |
| Open a sam/bam file for writing with the specified filename, determining SAM/BAM from the extension (.bam = BAM). | |
| bool | ReadBamIndex (const char *filename) |
| Read the specified bam index file. | |
| bool | ReadBamIndex () |
| Read the bam index file using the BAM filename as a base. | |
| void | SetReference (GenomeSequence *reference) |
| Sets the reference to the specified genome sequence object. | |
| void | SetReadSequenceTranslation (SamRecord::SequenceTranslation translation) |
| Set the type of sequence translation to use when reading the sequence. | |
| void | SetWriteSequenceTranslation (SamRecord::SequenceTranslation translation) |
| Set the type of sequence translation to use when writing the sequence. | |
| void | Close () |
| Close the file if there is one open. | |
| bool | IsOpen () |
| Returns whether or not the file has been opened successfully. | |
| bool | IsEOF () |
| Returns whether or not the end of the file has been reached. | |
| bool | ReadHeader (SamFileHeader &header) |
| Reads the header section from the file and stores it in the passed in header. | |
| bool | WriteHeader (SamFileHeader &header) |
| Writes the specified header into the file. | |
| bool | ReadRecord (SamFileHeader &header, SamRecord &record) |
| Reads the next record from the file & stores it in the passed in record. | |
| bool | WriteRecord (SamFileHeader &header, SamRecord &record) |
| Writes the specified record into the file. | |
| void | setSortedValidation (SortedType sortType) |
| Set the flag to validate that the file is sorted as it is read/written. | |
| uint32_t | GetCurrentRecordCount () |
| Return the number of records that have been read/written so far. | |
| SamStatus::Status | GetFailure () |
| Deprecated, get the Status of the last call that sets status. | |
| SamStatus::Status | GetStatus () |
| Get the Status of the last call that sets status. | |
| const char * | GetStatusMessage () |
| Get the Status Message of the last call that sets status. | |
| bool | SetReadSection (int32_t refID) |
| Sets which reference id (index into the BAM list of reference information) of the BAM file should be read. | |
| bool | SetReadSection (const char *refName) |
| Sets which reference name of the BAM file should be read. | |
| bool | SetReadSection (int32_t refID, int32_t start, int32_t end, bool overlap=true) |
| Sets which reference id (index into the BAM list of reference information) & start/end positions of the BAM file should be read. | |
| bool | SetReadSection (const char *refName, int32_t start, int32_t end, bool overlap=true) |
| Sets which reference name & start/end positions of the BAM file should be read. | |
| void | SetReadFlags (uint16_t requiredFlags, uint16_t excludedFlags) |
| Specify which reads should be returned by ReadRecord. | |
| int32_t | getNumMappedReadsFromIndex (int32_t refID) |
| Get the number of mapped reads in the specified reference id. | |
| int32_t | getNumUnMappedReadsFromIndex (int32_t refID) |
| Get the number of unmapped reads in the specified reference id. | |
| int32_t | getNumMappedReadsFromIndex (const char *refName, SamFileHeader &header) |
| Get the number of mapped reads in the specified reference name. | |
| int32_t | getNumUnMappedReadsFromIndex (const char *refName, SamFileHeader &header) |
| Get the number of unmapped reads in the specified reference name. | |
| uint32_t | GetNumOverlaps (SamRecord &samRecord) |
| Returns the number of bases in the passed in read that overlap the region that is currently set. | |
| void | GenerateStatistics (bool genStats) |
| Whether or not statistics should be generated for this file. | |
| const BamIndex * | GetBamIndex () |
| Return the bam index if one has been opened. | |
| long int | GetCurrentPosition () |
| Get the current file position. | |
| void | DisableBuffering () |
| Turn off file read buffering. | |
| void | PrintStatistics () |
| Print the statistics that have been recorded due to a call to GenerateStatistics. | |
| bool | attemptRecoverySync (bool(*checkSignature)(void *data), int length) |
| void | setAttemptRecovery (bool flag=false) |
Protected Member Functions | |
| void | init () |
| void | init (const char *filename, OpenType mode, SamFileHeader *header) |
| void | resetFile () |
| Resets the file prepping for a new file. | |
| bool | validateSortOrder (SamRecord &record, SamFileHeader &header) |
| Validate that the record is sorted compared to the previously read record if there is one, according to the specified sort order. | |
| SortedType | getSortOrderFromHeader (SamFileHeader &header) |
| bool | processNewSection (SamFileHeader &header) |
| bool | ensureIndexedReadPosition () |
| bool | checkRecordInSection (SamRecord &record) |
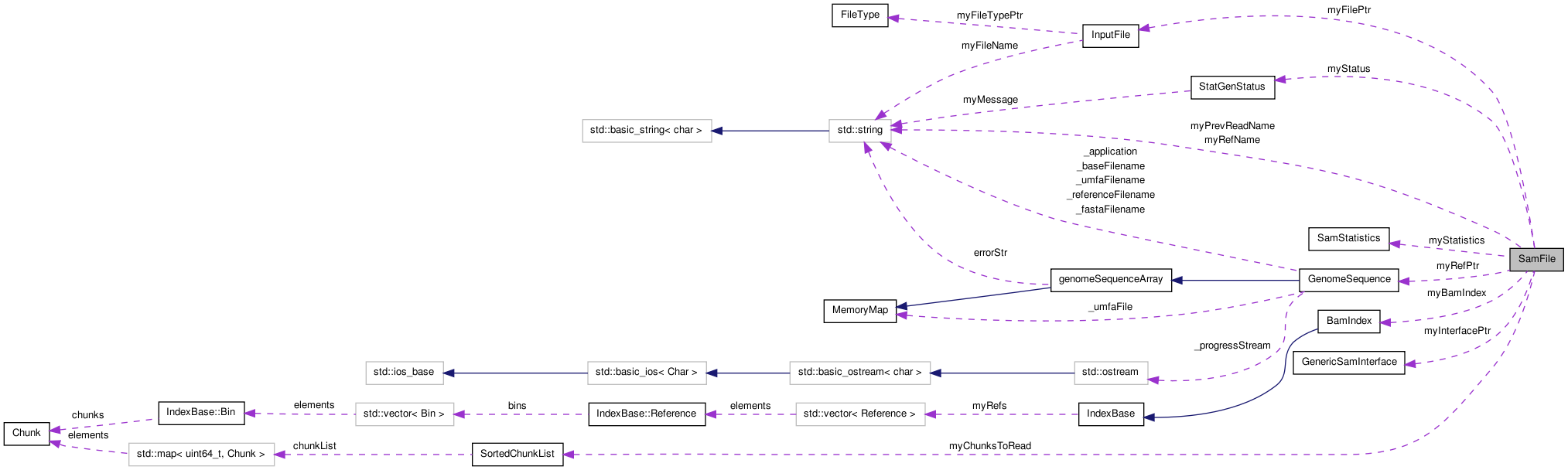
Protected Attributes | |
| IFILE | myFilePtr |
| GenericSamInterface * | myInterfacePtr |
| bool | myIsOpenForRead |
| Flag to indicate if a file is open for reading. | |
| bool | myIsOpenForWrite |
| Flag to indicate if a file is open for writing. | |
| bool | myHasHeader |
| Flag to indicate if a header has been read/written - required before being able to read/write a record. | |
| SortedType | mySortedType |
| int32_t | myPrevCoord |
| Previous values used for checking if the file is sorted. | |
| int32_t | myPrevRefID |
| std::string | myPrevReadName |
| uint32_t | myRecordCount |
| Keep a count of the number of records that have been read/written so far. | |
| SamStatistics * | myStatistics |
| Pointer to the statistics for this file. | |
| SamStatus | myStatus |
| The status of the last SamFile command. | |
| bool | myIsBamOpenForRead |
| Values for reading Sorted BAM files via the index. | |
| bool | myNewSection |
| bool | myOverlapSection |
| int32_t | myRefID |
| int32_t | myStartPos |
| int32_t | myEndPos |
| uint64_t | myCurrentChunkEnd |
| SortedChunkList | myChunksToRead |
| BamIndex * | myBamIndex |
| GenomeSequence * | myRefPtr |
| SamRecord::SequenceTranslation | myReadTranslation |
| SamRecord::SequenceTranslation | myWriteTranslation |
| std::string | myRefName |
Detailed Description
Allows the user to easily read/write a SAM/BAM file.
The SamFile class contains additional functionality that allows a user to read specific sections of sorted & indexed BAM files. In order to take advantage of this capability, the index file must be read prior to setting the read section. This logic saves the time of having to read the entire file and takes advantage of the seeking capability of BGZF.
Definition at line 35 of file SamFile.h.
Member Enumeration Documentation
| enum SamFile::OpenType |
| enum SamFile::SortedType |
Enum for indicating the type of sort expected in the file.
- Enumerator:
UNSORTED file is not sorted.
FLAG SO flag from the header indicates the sort type.
COORDINATE file is sorted by coordinate.
QUERY_NAME file is sorted by queryname.
Definition at line 46 of file SamFile.h.
00046 { 00047 UNSORTED = 0, ///< file is not sorted. 00048 FLAG, ///< SO flag from the header indicates the sort type. 00049 COORDINATE, ///< file is sorted by coordinate. 00050 QUERY_NAME ///< file is sorted by queryname. 00051 };
Constructor & Destructor Documentation
| SamFile::SamFile | ( | ) |
Default Constructor, initializes the variables, but does not open any files.
Definition at line 26 of file SamFile.cpp.
References resetFile().
| SamFile::SamFile | ( | ErrorHandler::HandlingType | errorHandlingType | ) |
Constructor that sets the error handling type.
- Parameters:
-
errorHandlingType how to handle errors.
Definition at line 35 of file SamFile.cpp.
References resetFile().
| SamFile::SamFile | ( | const char * | filename, | |
| OpenType | mode | |||
| ) |
Constructor that opens the specified file based on the specified mode (READ/WRITE), aborts if the file could not be opened.
- Parameters:
-
filename name of the file to open. mode mode to use for opening the file.
Definition at line 45 of file SamFile.cpp.
00046 : myStatus() 00047 { 00048 init(filename, mode, NULL); 00049 }
| SamFile::SamFile | ( | const char * | filename, | |
| OpenType | mode, | |||
| ErrorHandler::HandlingType | errorHandlingType | |||
| ) |
Constructor that opens the specified file based on the specified mode (READ/WRITE) and handles errors per the specified handleType.
- Parameters:
-
filename name of the file to open. mode mode to use for opening the file. errorHandlingType how to handle errors.
Definition at line 54 of file SamFile.cpp.
00056 : myStatus(errorHandlingType) 00057 { 00058 init(filename, mode, NULL); 00059 }
| SamFile::SamFile | ( | const char * | filename, | |
| OpenType | mode, | |||
| SamFileHeader * | header | |||
| ) |
Constructor that opens the specified file based on the specified mode (READ/WRITE) and reads the header, aborts if the file could not be opened or the header not read.
- Parameters:
-
filename name of the file to open. mode mode to use for opening the file. header to read into or write from
Definition at line 64 of file SamFile.cpp.
00065 : myStatus() 00066 { 00067 init(filename, mode, header); 00068 }
| SamFile::SamFile | ( | const char * | filename, | |
| OpenType | mode, | |||
| ErrorHandler::HandlingType | errorHandlingType, | |||
| SamFileHeader * | header | |||
| ) |
Constructor that opens the specified file based on the specified mode (READ/WRITE) and reads the header, handling errors per the specified handleType.
- Parameters:
-
filename name of the file to open. mode mode to use for opening the file. errorHandlingType how to handle errors. header to read into or write from
Definition at line 73 of file SamFile.cpp.
00076 : myStatus(errorHandlingType) 00077 { 00078 init(filename, mode, header); 00079 }
Member Function Documentation
| void SamFile::GenerateStatistics | ( | bool | genStats | ) |
Whether or not statistics should be generated for this file.
The value is carried over between files and is not reset, but the statistics themselves are reset between files.
- Parameters:
-
genStats set to true if statistics should be generated, false if not.
Definition at line 868 of file SamFile.cpp.
References myStatistics.
00869 { 00870 if(genStats) 00871 { 00872 if(myStatistics == NULL) 00873 { 00874 // Want to generate statistics, but do not yet have the 00875 // structure for them, so create one. 00876 myStatistics = new SamStatistics(); 00877 } 00878 } 00879 else 00880 { 00881 // Do not generate statistics, so if myStatistics is not NULL, 00882 // delete it. 00883 if(myStatistics != NULL) 00884 { 00885 delete myStatistics; 00886 myStatistics = NULL; 00887 } 00888 } 00889 00890 }
| const BamIndex * SamFile::GetBamIndex | ( | ) |
Return the bam index if one has been opened.
- Returns:
- const pointer to the bam index, or null if one has not been opened.
Definition at line 893 of file SamFile.cpp.
| long int SamFile::GetCurrentPosition | ( | ) | [inline] |
| SamStatus::Status SamFile::GetFailure | ( | ) | [inline] |
Deprecated, get the Status of the last call that sets status.
To remain backwards compatable - will be removed later.
Definition at line 201 of file SamFile.h.
References GetStatus().
00202 { 00203 return(GetStatus()); 00204 }
| int32_t SamFile::getNumMappedReadsFromIndex | ( | const char * | refName, | |
| SamFileHeader & | header | |||
| ) |
Get the number of mapped reads in the specified reference name.
Returns -1 for unknown reference names.
- Parameters:
-
refName reference name for which to extract the number of mapped reads. header header object containing the map from refName to refID
- Returns:
- number of mapped reads for the specified reference name.
Definition at line 810 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, BamIndex::getNumMappedReads(), SamFileHeader::getReferenceID(), myStatus, BamIndex::REF_ID_UNMAPPED, and StatGenStatus::setStatus().
00812 { 00813 // The bam index must have already been read. 00814 if(myBamIndex == NULL) 00815 { 00816 myStatus.setStatus(SamStatus::FAIL_ORDER, 00817 "Cannot get num mapped reads from the index until it has been read."); 00818 return(false); 00819 } 00820 int32_t refID = BamIndex::REF_ID_UNMAPPED; 00821 if((strcmp(refName, "") != 0) && (strcmp(refName, "*") != 0)) 00822 { 00823 // Reference name specified, so read just the "-1" entries. 00824 refID = header.getReferenceID(refName); 00825 } 00826 return(myBamIndex->getNumMappedReads(refID)); 00827 }
| int32_t SamFile::getNumMappedReadsFromIndex | ( | int32_t | refID | ) |
Get the number of mapped reads in the specified reference id.
Returns -1 for out of range refIDs.
- Parameters:
-
refID reference ID for which to extract the number of mapped reads.
- Returns:
- number of mapped reads for the specified reference id.
Definition at line 780 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, BamIndex::getNumMappedReads(), myStatus, and StatGenStatus::setStatus().
00781 { 00782 // The bam index must have already been read. 00783 if(myBamIndex == NULL) 00784 { 00785 myStatus.setStatus(SamStatus::FAIL_ORDER, 00786 "Cannot get num mapped reads from the index until it has been read."); 00787 return(false); 00788 } 00789 return(myBamIndex->getNumMappedReads(refID)); 00790 }
| uint32_t SamFile::GetNumOverlaps | ( | SamRecord & | samRecord | ) |
Returns the number of bases in the passed in read that overlap the region that is currently set.
Overlapping means that the bases occur in both the read and the reference as either matches or mismatches. This does not count insertions, deletions, clips, pads, or skips.
- Parameters:
-
samRecord to check for overlapping bases.
- Returns:
- number of bases that overlap region that is currently set.
Definition at line 854 of file SamFile.cpp.
References SamRecord::getNumOverlaps(), SamRecord::setReference(), and SamRecord::setSequenceTranslation().
00855 { 00856 if(myRefPtr != NULL) 00857 { 00858 samRecord.setReference(myRefPtr); 00859 } 00860 samRecord.setSequenceTranslation(myReadTranslation); 00861 00862 // Get the overlaps in the sam record for the region currently set 00863 // for this file. 00864 return(samRecord.getNumOverlaps(myStartPos, myEndPos)); 00865 }
| int32_t SamFile::getNumUnMappedReadsFromIndex | ( | const char * | refName, | |
| SamFileHeader & | header | |||
| ) |
Get the number of unmapped reads in the specified reference name.
Returns -1 for unknown reference names.
- Parameters:
-
refName reference name for which to extract the number of unmapped reads. header header object containing the map from refName to refID
- Returns:
- number of unmapped reads for the specified reference name.
Definition at line 832 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, BamIndex::getNumUnMappedReads(), SamFileHeader::getReferenceID(), myStatus, BamIndex::REF_ID_UNMAPPED, and StatGenStatus::setStatus().
00834 { 00835 // The bam index must have already been read. 00836 if(myBamIndex == NULL) 00837 { 00838 myStatus.setStatus(SamStatus::FAIL_ORDER, 00839 "Cannot get num unmapped reads from the index until it has been read."); 00840 return(false); 00841 } 00842 int32_t refID = BamIndex::REF_ID_UNMAPPED; 00843 if((strcmp(refName, "") != 0) && (strcmp(refName, "*") != 0)) 00844 { 00845 // Reference name specified, so read just the "-1" entries. 00846 refID = header.getReferenceID(refName); 00847 } 00848 return(myBamIndex->getNumUnMappedReads(refID)); 00849 }
| int32_t SamFile::getNumUnMappedReadsFromIndex | ( | int32_t | refID | ) |
Get the number of unmapped reads in the specified reference id.
Returns -1 for out of range refIDs.
- Parameters:
-
refID reference ID for which to extract the number of unmapped reads.
- Returns:
- number of unmapped reads for the specified reference id.
Definition at line 795 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, BamIndex::getNumUnMappedReads(), myStatus, and StatGenStatus::setStatus().
00796 { 00797 // The bam index must have already been read. 00798 if(myBamIndex == NULL) 00799 { 00800 myStatus.setStatus(SamStatus::FAIL_ORDER, 00801 "Cannot get num unmapped reads from the index until it has been read."); 00802 return(false); 00803 } 00804 return(myBamIndex->getNumUnMappedReads(refID)); 00805 }
| bool SamFile::IsEOF | ( | ) |
Returns whether or not the end of the file has been reached.
- Returns:
- true = EOF; false = not eof. If the file is not open, true is returned.
Definition at line 414 of file SamFile.cpp.
References ifeof().
00415 { 00416 if (myFilePtr != NULL) 00417 { 00418 // File Pointer is set, so return if eof. 00419 return(ifeof(myFilePtr)); 00420 } 00421 // File pointer is not set, so return true, eof. 00422 return true; 00423 }
| bool SamFile::IsOpen | ( | ) |
Returns whether or not the file has been opened successfully.
- Returns:
- true = open; false = not open.
Definition at line 400 of file SamFile.cpp.
References InputFile::isOpen().
00401 { 00402 if (myFilePtr != NULL) 00403 { 00404 // File Pointer is set, so return if it is open. 00405 return(myFilePtr->isOpen()); 00406 } 00407 // File pointer is not set, so return false, not open. 00408 return false; 00409 }
| bool SamFile::OpenForRead | ( | const char * | filename, | |
| SamFileHeader * | header = NULL | |||
| ) |
Open a sam/bam file for reading with the specified filename, determing the type of file and SAM/BAM by reading the file (if not stdin).
- Parameters:
-
filename the sam/bam file to open for reading. header to read into or write from (optional)
- Returns:
- true = success; false = failure.
Definition at line 93 of file SamFile.cpp.
References InputFile::BGZF, InputFile::DEFAULT, StatGenStatus::FAIL_IO, ifopen(), ifread(), ifrewind(), myIsBamOpenForRead, myIsOpenForRead, myStatus, ReadHeader(), resetFile(), InputFile::setAttemptRecovery(), StatGenStatus::setStatus(), StatGenStatus::SUCCESS, and InputFile::UNCOMPRESSED.
Referenced by Pileup< PILEUP_TYPE, FUNC_CLASS >::processFile().
00094 { 00095 // Reset for any previously operated on files. 00096 resetFile(); 00097 00098 int lastchar = 0; 00099 00100 while (filename[lastchar] != 0) lastchar++; 00101 00102 // If at least one character, check for '-'. 00103 if((lastchar >= 1) && (filename[0] == '-')) 00104 { 00105 // Read from stdin - determine type of file to read. 00106 // Determine if compressed bam. 00107 if(strcmp(filename, "-.bam") == 0) 00108 { 00109 // Compressed bam - open as bgzf. 00110 // -.bam is the filename, read compressed bam from stdin 00111 filename = "-"; 00112 00113 myFilePtr = new InputFile; 00114 // support recover mode - this switches in a reader 00115 // capable of recovering from bad BGZF compression blocks. 00116 myFilePtr->setAttemptRecovery(myAttemptRecovery); 00117 myFilePtr->openFile(filename, "rb", InputFile::BGZF); 00118 00119 myInterfacePtr = new BamInterface; 00120 00121 // Read the magic string. 00122 char magic[4]; 00123 ifread(myFilePtr, magic, 4); 00124 } 00125 else if(strcmp(filename, "-.ubam") == 0) 00126 { 00127 // uncompressed BAM File. 00128 // -.ubam is the filename, read uncompressed bam from stdin. 00129 // uncompressed BAM is still compressed with BGZF, but using 00130 // compression level 0, so still open as BGZF since it has a 00131 // BGZF header. 00132 filename = "-"; 00133 00134 // Uncompressed, so do not require the eof block. 00135 #ifdef __ZLIB_AVAILABLE__ 00136 BgzfFileType::setRequireEofBlock(false); 00137 #endif 00138 myFilePtr = ifopen(filename, "rb", InputFile::BGZF); 00139 00140 myInterfacePtr = new BamInterface; 00141 00142 // Read the magic string. 00143 char magic[4]; 00144 ifread(myFilePtr, magic, 4); 00145 } 00146 else 00147 { 00148 // SAM File. 00149 // read sam from stdin 00150 filename = "-"; 00151 myFilePtr = ifopen(filename, "rb", InputFile::UNCOMPRESSED); 00152 myInterfacePtr = new SamInterface; 00153 } 00154 } 00155 else 00156 { 00157 // Not from stdin. Read the file to determine the type. 00158 00159 myFilePtr = new InputFile; 00160 00161 // support recovery mode - this conditionally enables a reader 00162 // capable of recovering from bad BGZF compression blocks. 00163 myFilePtr->setAttemptRecovery(myAttemptRecovery); 00164 bool rc = myFilePtr->openFile(filename, "rb", InputFile::DEFAULT); 00165 00166 if (rc == false) 00167 { 00168 std::string errorMessage = "Failed to Open "; 00169 errorMessage += filename; 00170 errorMessage += " for reading"; 00171 myStatus.setStatus(SamStatus::FAIL_IO, errorMessage.c_str()); 00172 delete myFilePtr; 00173 myFilePtr = NULL; 00174 return(false); 00175 } 00176 00177 char magic[4]; 00178 ifread(myFilePtr, magic, 4); 00179 00180 if (magic[0] == 'B' && magic[1] == 'A' && magic[2] == 'M' && 00181 magic[3] == 1) 00182 { 00183 myInterfacePtr = new BamInterface; 00184 // Set that it is a bam file open for reading. This is needed to 00185 // determine if an index file can be used. 00186 myIsBamOpenForRead = true; 00187 } 00188 else 00189 { 00190 // Not a bam, so rewind to the beginning of the file so it 00191 // can be read. 00192 ifrewind(myFilePtr); 00193 myInterfacePtr = new SamInterface; 00194 } 00195 } 00196 00197 // File is open for reading. 00198 myIsOpenForRead = true; 00199 00200 // Read the header if one was passed in. 00201 if(header != NULL) 00202 { 00203 return(ReadHeader(*header)); 00204 } 00205 00206 // Successfully opened the file. 00207 myStatus = SamStatus::SUCCESS; 00208 return(true); 00209 }
| bool SamFile::OpenForWrite | ( | const char * | filename, | |
| SamFileHeader * | header = NULL | |||
| ) |
Open a sam/bam file for writing with the specified filename, determining SAM/BAM from the extension (.bam = BAM).
- Parameters:
-
filename the sam/bam file to open for writing. header to read into or write from (optional)
- Returns:
- true = success; false = failure.
Definition at line 213 of file SamFile.cpp.
References InputFile::BGZF, StatGenStatus::FAIL_IO, ifopen(), myIsOpenForWrite, myStatus, resetFile(), StatGenStatus::setStatus(), StatGenStatus::SUCCESS, InputFile::UNCOMPRESSED, and WriteHeader().
00214 { 00215 // Reset for any previously operated on files. 00216 resetFile(); 00217 00218 int lastchar = 0; 00219 while (filename[lastchar] != 0) lastchar++; 00220 if (lastchar >= 4 && 00221 filename[lastchar - 4] == 'u' && 00222 filename[lastchar - 3] == 'b' && 00223 filename[lastchar - 2] == 'a' && 00224 filename[lastchar - 1] == 'm') 00225 { 00226 // BAM File. 00227 // if -.ubam is the filename, write uncompressed bam to stdout 00228 if((lastchar == 6) && (filename[0] == '-') && (filename[1] == '.')) 00229 { 00230 filename = "-"; 00231 } 00232 00233 myFilePtr = ifopen(filename, "wb0", InputFile::BGZF); 00234 00235 myInterfacePtr = new BamInterface; 00236 } 00237 else if (lastchar >= 3 && 00238 filename[lastchar - 3] == 'b' && 00239 filename[lastchar - 2] == 'a' && 00240 filename[lastchar - 1] == 'm') 00241 { 00242 // BAM File. 00243 // if -.bam is the filename, write compressed bam to stdout 00244 if((lastchar == 5) && (filename[0] == '-') && (filename[1] == '.')) 00245 { 00246 filename = "-"; 00247 } 00248 myFilePtr = ifopen(filename, "wb", InputFile::BGZF); 00249 00250 myInterfacePtr = new BamInterface; 00251 } 00252 else 00253 { 00254 // SAM File 00255 // if - (followed by anything is the filename, 00256 // write uncompressed sam to stdout 00257 if((lastchar >= 1) && (filename[0] == '-')) 00258 { 00259 filename = "-"; 00260 } 00261 myFilePtr = ifopen(filename, "wb", InputFile::UNCOMPRESSED); 00262 00263 myInterfacePtr = new SamInterface; 00264 } 00265 00266 if (myFilePtr == NULL) 00267 { 00268 std::string errorMessage = "Failed to Open "; 00269 errorMessage += filename; 00270 errorMessage += " for writing"; 00271 myStatus.setStatus(SamStatus::FAIL_IO, errorMessage.c_str()); 00272 return(false); 00273 } 00274 00275 myIsOpenForWrite = true; 00276 00277 // Write the header if one was passed in. 00278 if(header != NULL) 00279 { 00280 return(WriteHeader(*header)); 00281 } 00282 00283 // Successfully opened the file. 00284 myStatus = SamStatus::SUCCESS; 00285 return(true); 00286 }
| void SamFile::PrintStatistics | ( | ) | [inline] |
Print the statistics that have been recorded due to a call to GenerateStatistics.
Definition at line 352 of file SamFile.h.
References myStatistics.
00352 {if(myStatistics != NULL) myStatistics->print();}
| bool SamFile::ReadBamIndex | ( | ) |
Read the bam index file using the BAM filename as a base.
It must be read prior to setting a read section, for seeking and reading portions of a bam file. Must be read after opening the BAM file since it uses the BAM filename as a base name for the index file. First it tries filename.bam.bai. If that fails, it tries it without the .bam extension, filename.bai.
- Returns:
- true = success; false = failure.
Definition at line 318 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, InputFile::getFileName(), myStatus, and StatGenStatus::setStatus().
00319 { 00320 if(myFilePtr == NULL) 00321 { 00322 // Can't read the bam index file because the BAM file has not yet been 00323 // opened, so we don't know the base filename for the index file. 00324 std::string errorMessage = "Failed to read the bam Index file -" 00325 " the BAM file needs to be read first in order to determine" 00326 " the index filename."; 00327 myStatus.setStatus(SamStatus::FAIL_ORDER, errorMessage.c_str()); 00328 return(false); 00329 } 00330 00331 const char* bamBaseName = myFilePtr->getFileName(); 00332 00333 std::string indexName = bamBaseName; 00334 indexName += ".bai"; 00335 00336 bool foundFile = true; 00337 try 00338 { 00339 if(ReadBamIndex(indexName.c_str()) == false) 00340 { 00341 foundFile = false; 00342 } 00343 } 00344 catch (std::exception& e) 00345 { 00346 foundFile = false; 00347 } 00348 00349 // Check to see if the index file was found. 00350 if(!foundFile) 00351 { 00352 // Not found - try without the bam extension. 00353 // Locate the start of the bam extension 00354 size_t startExt = indexName.find(".bam"); 00355 if(startExt == std::string::npos) 00356 { 00357 // Could not find the .bam extension, so just return false since the 00358 // call to ReadBamIndex set the status. 00359 return(false); 00360 } 00361 // Remove ".bam" and try reading the index again. 00362 indexName.erase(startExt, 4); 00363 return(ReadBamIndex(indexName.c_str())); 00364 } 00365 return(true); 00366 }
| bool SamFile::ReadBamIndex | ( | const char * | filename | ) |
Read the specified bam index file.
It must be read prior to setting a read section, for seeking and reading portions of a bam file.
- Parameters:
-
filename the name of the bam index file to be read.
- Returns:
- true = success; false = failure.
Definition at line 290 of file SamFile.cpp.
References myStatus, BamIndex::readIndex(), StatGenStatus::setStatus(), and StatGenStatus::SUCCESS.
00291 { 00292 // Cleanup a previously setup index. 00293 if(myBamIndex != NULL) 00294 { 00295 delete myBamIndex; 00296 myBamIndex = NULL; 00297 } 00298 00299 // Create a new bam index. 00300 myBamIndex = new BamIndex(); 00301 SamStatus::Status indexStat = myBamIndex->readIndex(bamIndexFilename); 00302 00303 if(indexStat != SamStatus::SUCCESS) 00304 { 00305 std::string errorMessage = "Failed to read the bam Index file: "; 00306 errorMessage += bamIndexFilename; 00307 myStatus.setStatus(indexStat, errorMessage.c_str()); 00308 delete myBamIndex; 00309 myBamIndex = NULL; 00310 return(false); 00311 } 00312 myStatus = SamStatus::SUCCESS; 00313 return(true); 00314 }
| bool SamFile::ReadHeader | ( | SamFileHeader & | header | ) |
Reads the header section from the file and stores it in the passed in header.
- Returns:
- true = success; false = failure.
Definition at line 427 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, myHasHeader, myIsOpenForRead, myStatus, StatGenStatus::setStatus(), and StatGenStatus::SUCCESS.
Referenced by OpenForRead(), and Pileup< PILEUP_TYPE, FUNC_CLASS >::processFile().
00428 { 00429 myStatus = SamStatus::SUCCESS; 00430 if(myIsOpenForRead == false) 00431 { 00432 // File is not open for read 00433 myStatus.setStatus(SamStatus::FAIL_ORDER, 00434 "Cannot read header since the file is not open for reading"); 00435 return(false); 00436 } 00437 00438 if(myHasHeader == true) 00439 { 00440 // The header has already been read. 00441 myStatus.setStatus(SamStatus::FAIL_ORDER, 00442 "Cannot read header since it has already been read."); 00443 return(false); 00444 } 00445 00446 if(myInterfacePtr->readHeader(myFilePtr, header, myStatus)) 00447 { 00448 // The header has now been successfully read. 00449 myHasHeader = true; 00450 return(true); 00451 } 00452 return(false); 00453 }
| bool SamFile::ReadRecord | ( | SamFileHeader & | header, | |
| SamRecord & | record | |||
| ) |
Reads the next record from the file & stores it in the passed in record.
If it is an indexed BAM file and SetReadSection was called, only alignments in the section specified by SetReadSection are read. If they all have already been read, this method returns false.
Validates that the record is sorted according to the value set by setSortedValidation. No sorting validation is done if specified to be unsorted, or setSortedValidation was never called.
- Returns:
- true = record was successfully set (and sorted if applicable), false = record was not successfully set (or not sorted as expected).
Definition at line 491 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, SamRecord::getFlag(), myHasHeader, myIsOpenForRead, myRecordCount, myStatistics, myStatus, SamRecord::setReference(), SamRecord::setSequenceTranslation(), StatGenStatus::setStatus(), StatGenStatus::SUCCESS, and validateSortOrder().
Referenced by Pileup< PILEUP_TYPE, FUNC_CLASS >::processFile().
00493 { 00494 myStatus = SamStatus::SUCCESS; 00495 00496 if(myIsOpenForRead == false) 00497 { 00498 // File is not open for read 00499 myStatus.setStatus(SamStatus::FAIL_ORDER, 00500 "Cannot read record since the file is not open for reading"); 00501 throw(std::runtime_error("SOFTWARE BUG: trying to read a SAM/BAM record prior to opening the file.")); 00502 return(false); 00503 } 00504 00505 if(myHasHeader == false) 00506 { 00507 // The header has not yet been read. 00508 // TODO - maybe just read the header. 00509 myStatus.setStatus(SamStatus::FAIL_ORDER, 00510 "Cannot read record since the header has not been read."); 00511 throw(std::runtime_error("SOFTWARE BUG: trying to read a SAM/BAM record prior to reading the header.")); 00512 return(false); 00513 } 00514 00515 // Check to see if a new region has been set. If so, determine the 00516 // chunks for that region. 00517 if(myNewSection) 00518 { 00519 if(!processNewSection(header)) 00520 { 00521 // Failed processing a new section. Could be an 00522 // order issue like the file not being open or the 00523 // indexed file not having been read. 00524 // processNewSection sets myStatus with the failure reason. 00525 return(false); 00526 } 00527 } 00528 00529 // Read until a record is not successfully read or there are no more 00530 // requested records. 00531 while(myStatus == SamStatus::SUCCESS) 00532 { 00533 record.setReference(myRefPtr); 00534 record.setSequenceTranslation(myReadTranslation); 00535 00536 // If reading by index, this method will setup to ensure it is in 00537 // the correct position for the next record (if not already there). 00538 // Sets myStatus if it could not move to a good section. 00539 // Just returns true if it is not setup to read by index. 00540 if(!ensureIndexedReadPosition()) 00541 { 00542 // Either there are no more records in the section 00543 // or it failed to move to the right section, so there 00544 // is nothing more to read, stop looping. 00545 break; 00546 } 00547 00548 // File is open for reading and the header has been read, so read the 00549 // next record. 00550 myInterfacePtr->readRecord(myFilePtr, header, record, myStatus); 00551 if(myStatus != SamStatus::SUCCESS) 00552 { 00553 // Failed to read the record, so break out of the loop. 00554 break; 00555 } 00556 00557 // Successfully read a record, so check if we should filter it. 00558 // First check if it is out of the section. Returns true 00559 // if not reading by sections, returns false if the record 00560 // is outside of the section. Sets status to NO_MORE_RECS if 00561 // there is nothing left ot read in the section. 00562 if(!checkRecordInSection(record)) 00563 { 00564 // The record is not in the section. 00565 // The while loop will detect if NO_MORE_RECS was set. 00566 continue; 00567 } 00568 00569 // Check the flag for required/excluded flags. 00570 uint16_t flag = record.getFlag(); 00571 if((flag & myRequiredFlags) != myRequiredFlags) 00572 { 00573 // The record does not conatain all required flags, so 00574 // continue looking. 00575 continue; 00576 } 00577 if((flag & myExcludedFlags) != 0) 00578 { 00579 // The record contains an excluded flag, so continue looking. 00580 continue; 00581 } 00582 00583 //increment the record count. 00584 myRecordCount++; 00585 00586 if(myStatistics != NULL) 00587 { 00588 // Statistics should be updated. 00589 myStatistics->updateStatistics(record); 00590 } 00591 00592 // Successfully read the record, so check the sort order. 00593 if(!validateSortOrder(record, header)) 00594 { 00595 // ValidateSortOrder sets the status on a failure. 00596 return(false); 00597 } 00598 return(true); 00599 00600 } // End while loop that checks if a desired record is found or failure. 00601 00602 // Return true if a record was found. 00603 return(myStatus == SamStatus::SUCCESS); 00604 }
| void SamFile::SetReadFlags | ( | uint16_t | requiredFlags, | |
| uint16_t | excludedFlags | |||
| ) |
Specify which reads should be returned by ReadRecord.
Reads will only be returned by ReadRecord that contain the specified required flags and that do not contain any of the specified excluded flags. ReadRecord will continue to read from the file until a record that complies with these flag settings is found or until the end of the file/region.
- Parameters:
-
requiredFlags flags that are required to be in records returned by ReadRecord (set to 0x0 if there are no required flags). excludedFlags flags that are required to not be in records returned by ReadRecord (set to 0x0 if there are no excluded flags).
Definition at line 771 of file SamFile.cpp.
| bool SamFile::SetReadSection | ( | const char * | refName, | |
| int32_t | start, | |||
| int32_t | end, | |||
| bool | overlap = true | |||
| ) |
Sets which reference name & start/end positions of the BAM file should be read.
The records for this reference name & positions will be retrieved on each ReadRecord call. Specify "" or "*" to indicate reads with no reference. When all records have been retrieved for the specified section, ReadRecord will return failure until a new read section is set. Must be called only after the file has been opened for reading. Sorting validation is reset everytime SetReadSection is called since it can jump around in the file.
- Parameters:
-
refName the reference name of the records to read from the file. start inclusive 0-based start position of records that should be read for this refID. end exclusive 0-based end position of records that should be read for this refID. overlap When true (default), return reads that just overlap the region; when false, only return reads that fall completely within the region
- Returns:
- true = success; false = failure.
Definition at line 726 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, myIsBamOpenForRead, myPrevCoord, myStatus, BamIndex::REF_ID_ALL, BamIndex::REF_ID_UNMAPPED, StatGenStatus::setStatus(), and StatGenStatus::SUCCESS.
00728 { 00729 // If there is not a BAM file open for reading, return failure. 00730 // Opening a new file clears the read section, so it must be 00731 // set after the file is opened. 00732 if(!myIsBamOpenForRead) 00733 { 00734 // There is not a BAM file open for reading. 00735 myStatus.setStatus(SamStatus::FAIL_ORDER, 00736 "Cannot set section since there is no bam file open"); 00737 return(false); 00738 } 00739 00740 myNewSection = true; 00741 myOverlapSection = overlap; 00742 myStartPos = start; 00743 myEndPos = end; 00744 if((strcmp(refName, "") == 0) || (strcmp(refName, "*") == 0)) 00745 { 00746 // No Reference name specified, so read just the "-1" entries. 00747 myRefID = BamIndex::REF_ID_UNMAPPED; 00748 } 00749 else 00750 { 00751 // save the reference name and revert the reference ID to unknown 00752 // so it will be calculated later. 00753 myRefName = refName; 00754 myRefID = BamIndex::REF_ID_ALL; 00755 } 00756 myChunksToRead.clear(); 00757 // Reset the end of the current chunk. We are resetting our read, so 00758 // we no longer have a "current chunk" that we are reading. 00759 myCurrentChunkEnd = 0; 00760 myStatus = SamStatus::SUCCESS; 00761 00762 // Reset the sort order criteria since we moved around in the file. 00763 myPrevCoord = -1; 00764 myPrevRefID = 0; 00765 myPrevReadName.clear(); 00766 00767 return(true); 00768 }
| bool SamFile::SetReadSection | ( | int32_t | refID, | |
| int32_t | start, | |||
| int32_t | end, | |||
| bool | overlap = true | |||
| ) |
Sets which reference id (index into the BAM list of reference information) & start/end positions of the BAM file should be read.
The records for that reference id and positions will be retrieved on each ReadRecord call. Reference ids start at 0, and -1 indicates reads with no reference. When all records have been retrieved for the specified reference id, ReadRecord will return failure until a new read section is set. Must be called only after the file has been opened for reading. Sorting validation is reset everytime SetReadPosition is called since it can jump around in the file.
- Parameters:
-
refID the reference ID of the records to read from the file. start inclusive 0-based start position of records that should be read for this refID. end exclusive 0-based end position of records that should be read for this refID. overlap When true (default), return reads that just overlap the region; when false, only return reads that fall completely within the region
- Returns:
- true = success; false = failure.
Definition at line 690 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, myIsBamOpenForRead, myPrevCoord, myStatus, StatGenStatus::setStatus(), and StatGenStatus::SUCCESS.
00692 { 00693 // If there is not a BAM file open for reading, return failure. 00694 // Opening a new file clears the read section, so it must be 00695 // set after the file is opened. 00696 if(!myIsBamOpenForRead) 00697 { 00698 // There is not a BAM file open for reading. 00699 myStatus.setStatus(SamStatus::FAIL_ORDER, 00700 "Cannot set section since there is no bam file open"); 00701 return(false); 00702 } 00703 00704 myNewSection = true; 00705 myOverlapSection = overlap; 00706 myStartPos = start; 00707 myEndPos = end; 00708 myRefID = refID; 00709 myRefName.clear(); 00710 myChunksToRead.clear(); 00711 // Reset the end of the current chunk. We are resetting our read, so 00712 // we no longer have a "current chunk" that we are reading. 00713 myCurrentChunkEnd = 0; 00714 myStatus = SamStatus::SUCCESS; 00715 00716 // Reset the sort order criteria since we moved around in the file. 00717 myPrevCoord = -1; 00718 myPrevRefID = 0; 00719 myPrevReadName.clear(); 00720 00721 return(true); 00722 }
| bool SamFile::SetReadSection | ( | const char * | refName | ) |
Sets which reference name of the BAM file should be read.
The records for that reference name will be retrieved on each ReadRecord call. Specify "" or "*" to read records not associated with a reference. When all records have been retrieved for the specified reference name, ReadRecord will return failure until a new read section is set. Must be called only after the file has been opened for reading. Sorting validation is reset everytime SetReadPosition is called since it can jump around in the file.
- Parameters:
-
refName the reference name of the records to read from the file.
- Returns:
- true = success; false = failure.
Definition at line 682 of file SamFile.cpp.
References SetReadSection().
00683 { 00684 // No start/end specified, so set back to default -1. 00685 return(SetReadSection(refName, -1, -1)); 00686 }
| bool SamFile::SetReadSection | ( | int32_t | refID | ) |
Sets which reference id (index into the BAM list of reference information) of the BAM file should be read.
The records for that reference id will be retrieved on each ReadRecord call. Reference ids start at 0, and -1 indicates reads with no reference. When all records have been retrieved for the specified reference id, ReadRecord will return failure until a new read section is set. Must be called only after the file has been opened for reading. Sorting validation is reset everytime SetReadPosition is called since it can jump around in the file.
- Parameters:
-
refID the reference ID of the records to read from the file.
- Returns:
- true = success; false = failure.
Definition at line 673 of file SamFile.cpp.
Referenced by SetReadSection().
00674 { 00675 // No start/end specified, so set back to default -1. 00676 return(SetReadSection(refID, -1, -1)); 00677 }
| void SamFile::SetReadSequenceTranslation | ( | SamRecord::SequenceTranslation | translation | ) |
Set the type of sequence translation to use when reading the sequence.
Passed down to the SamRecord when it is read. The default type (if this method is never called) is NONE (the sequence is left as-is).
- Parameters:
-
translation type of sequence translation to use.
Definition at line 377 of file SamFile.cpp.
| void SamFile::SetReference | ( | GenomeSequence * | reference | ) |
Sets the reference to the specified genome sequence object.
- Parameters:
-
reference pointer to the GenomeSequence object.
Definition at line 370 of file SamFile.cpp.
Referenced by Pileup< PILEUP_TYPE, FUNC_CLASS >::processFile().
| void SamFile::setSortedValidation | ( | SortedType | sortType | ) |
Set the flag to validate that the file is sorted as it is read/written.
Must be called after the file has been opened. Sorting validation is reset everytime SetReadPosition is called since it can jump around in the file.
- Parameters:
-
sortType specifies the type of sort to be checked for.
Definition at line 659 of file SamFile.cpp.
Referenced by Pileup< PILEUP_TYPE, FUNC_CLASS >::processFile().
| void SamFile::SetWriteSequenceTranslation | ( | SamRecord::SequenceTranslation | translation | ) |
Set the type of sequence translation to use when writing the sequence.
Passed down to the SamRecord when it is written. The default type (if this method is never called) is NONE (the sequence is left as-is).
- Parameters:
-
translation type of sequence translation to use.
Definition at line 384 of file SamFile.cpp.
| bool SamFile::validateSortOrder | ( | SamRecord & | record, | |
| SamFileHeader & | header | |||
| ) | [protected] |
Validate that the record is sorted compared to the previously read record if there is one, according to the specified sort order.
If the sort order is UNSORTED, true is returned. Sorting validation is reset everytime SetReadPosition is called since it can jump around in the file.
Definition at line 996 of file SamFile.cpp.
References FLAG, SamRecord::get0BasedPosition(), SamRecord::getReadName(), SamRecord::getReferenceID(), StatGenStatus::INVALID_SORT, myPrevCoord, myRecordCount, myStatus, QUERY_NAME, BamIndex::REF_ID_UNMAPPED, SamRecord::setReference(), SamRecord::setSequenceTranslation(), StatGenStatus::setStatus(), and UNSORTED.
Referenced by ReadRecord(), and WriteRecord().
00997 { 00998 if(myRefPtr != NULL) 00999 { 01000 record.setReference(myRefPtr); 01001 } 01002 record.setSequenceTranslation(myReadTranslation); 01003 01004 bool status = false; 01005 if(mySortedType == UNSORTED) 01006 { 01007 // Unsorted, so nothing to validate, just return true. 01008 status = true; 01009 } 01010 else 01011 { 01012 // Check to see if mySortedType is based on the header. 01013 if(mySortedType == FLAG) 01014 { 01015 // Determine the sorted type from what was read out of the header. 01016 mySortedType = getSortOrderFromHeader(header); 01017 } 01018 01019 if(mySortedType == QUERY_NAME) 01020 { 01021 // Validate that it is sorted by query name. 01022 // Get the query name from the record. 01023 const char* readName = record.getReadName(); 01024 if(myPrevReadName.compare(readName) > 0) 01025 { 01026 // The previous name is greater than the new record's name, so 01027 // return false. 01028 String errorMessage = "ERROR: File is not sorted at record "; 01029 errorMessage += myRecordCount; 01030 myStatus.setStatus(SamStatus::INVALID_SORT, 01031 errorMessage.c_str()); 01032 status = false; 01033 } 01034 else 01035 { 01036 myPrevReadName = readName; 01037 status = true; 01038 } 01039 } 01040 else 01041 { 01042 // Validate that it is sorted by COORDINATES. 01043 // Get the leftmost coordinate and the reference index. 01044 int32_t refID = record.getReferenceID(); 01045 int32_t coord = record.get0BasedPosition(); 01046 // The unmapped reference id is at the end of a sorted file. 01047 if(refID == BamIndex::REF_ID_UNMAPPED) 01048 { 01049 // A new reference ID that is for the unmapped reads 01050 // is always valid. 01051 status = true; 01052 myPrevRefID = refID; 01053 myPrevCoord = coord; 01054 } 01055 else if(myPrevRefID == BamIndex::REF_ID_UNMAPPED) 01056 { 01057 // Previous reference ID was for unmapped reads, but the 01058 // current one is not, so this is not sorted. 01059 String errorMessage = "ERROR: File is not sorted at record "; 01060 errorMessage += myRecordCount; 01061 myStatus.setStatus(SamStatus::INVALID_SORT, 01062 errorMessage.c_str()); 01063 status = false; 01064 } 01065 else if(refID < myPrevRefID) 01066 { 01067 // Current reference id is less than the previous one, 01068 //meaning that it is not sorted. 01069 String errorMessage = "ERROR: File is not sorted at record "; 01070 errorMessage += myRecordCount; 01071 myStatus.setStatus(SamStatus::INVALID_SORT, 01072 errorMessage.c_str()); 01073 status = false; 01074 } 01075 else 01076 { 01077 // The reference IDs are in the correct order. 01078 if(refID > myPrevRefID) 01079 { 01080 // New reference id, so set the previous coordinate to -1 01081 myPrevCoord = -1; 01082 } 01083 01084 // Check the coordinates. 01085 if(coord < myPrevCoord) 01086 { 01087 // New Coord is less than the previous position. 01088 String errorMessage = "ERROR: File is not sorted at record "; 01089 errorMessage += myRecordCount; 01090 myStatus.setStatus(SamStatus::INVALID_SORT, 01091 errorMessage.c_str()); 01092 status = false; 01093 } 01094 else 01095 { 01096 myPrevRefID = refID; 01097 myPrevCoord = coord; 01098 status = true; 01099 } 01100 } 01101 } 01102 } 01103 01104 return(status); 01105 }
| bool SamFile::WriteHeader | ( | SamFileHeader & | header | ) |
Writes the specified header into the file.
- Returns:
- true = success; false = failure.
Definition at line 457 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, myHasHeader, myIsOpenForWrite, myStatus, StatGenStatus::setStatus(), and StatGenStatus::SUCCESS.
Referenced by OpenForWrite().
00458 { 00459 myStatus = SamStatus::SUCCESS; 00460 if(myIsOpenForWrite == false) 00461 { 00462 // File is not open for write 00463 // -OR- 00464 // The header has already been written. 00465 myStatus.setStatus(SamStatus::FAIL_ORDER, 00466 "Cannot write header since the file is not open for writing"); 00467 return(false); 00468 } 00469 00470 if(myHasHeader == true) 00471 { 00472 // The header has already been written. 00473 myStatus.setStatus(SamStatus::FAIL_ORDER, 00474 "Cannot write header since it has already been written"); 00475 return(false); 00476 } 00477 00478 if(myInterfacePtr->writeHeader(myFilePtr, header, myStatus)) 00479 { 00480 // The header has now been successfully written. 00481 myHasHeader = true; 00482 return(true); 00483 } 00484 00485 // return the status. 00486 return(false); 00487 }
| bool SamFile::WriteRecord | ( | SamFileHeader & | header, | |
| SamRecord & | record | |||
| ) |
Writes the specified record into the file.
Validates that the record is sorted according to the value set by setSortedValidation. No sorting validation is done if specified to be unsorted, or setSortedValidation was never called. Returns false and does not write the record if the record was not properly sorted.
- Returns:
- true = success; false = failure.
Definition at line 609 of file SamFile.cpp.
References StatGenStatus::FAIL_ORDER, StatGenStatus::INVALID_SORT, myHasHeader, myIsOpenForWrite, myRecordCount, myStatus, SamRecord::setReference(), StatGenStatus::setStatus(), StatGenStatus::SUCCESS, and validateSortOrder().
Referenced by SamCoordOutput::flush().
00611 { 00612 if(myIsOpenForWrite == false) 00613 { 00614 // File is not open for writing 00615 myStatus.setStatus(SamStatus::FAIL_ORDER, 00616 "Cannot write record since the file is not open for writing"); 00617 return(false); 00618 } 00619 00620 if(myHasHeader == false) 00621 { 00622 // The header has not yet been written. 00623 myStatus.setStatus(SamStatus::FAIL_ORDER, 00624 "Cannot write record since the header has not been written"); 00625 return(false); 00626 } 00627 00628 // Before trying to write the record, validate the sort order. 00629 if(!validateSortOrder(record, header)) 00630 { 00631 // Not sorted like it is supposed to be, do not write the record 00632 myStatus.setStatus(SamStatus::INVALID_SORT, 00633 "Cannot write the record since the file is not properly sorted."); 00634 return(false); 00635 } 00636 00637 if(myRefPtr != NULL) 00638 { 00639 record.setReference(myRefPtr); 00640 } 00641 00642 // File is open for writing and the header has been written, so write the 00643 // record. 00644 myStatus = myInterfacePtr->writeRecord(myFilePtr, header, record, 00645 myWriteTranslation); 00646 00647 if(myStatus == SamStatus::SUCCESS) 00648 { 00649 // A record was successfully written, so increment the record count. 00650 myRecordCount++; 00651 return(true); 00652 } 00653 return(false); 00654 }
Member Data Documentation
bool SamFile::myHasHeader [protected] |
Flag to indicate if a header has been read/written - required before being able to read/write a record.
Definition at line 399 of file SamFile.h.
Referenced by ReadHeader(), ReadRecord(), resetFile(), WriteHeader(), and WriteRecord().
The documentation for this class was generated from the following files:
- bam/SamFile.h
- bam/SamFile.cpp
 1.6.3
1.6.3